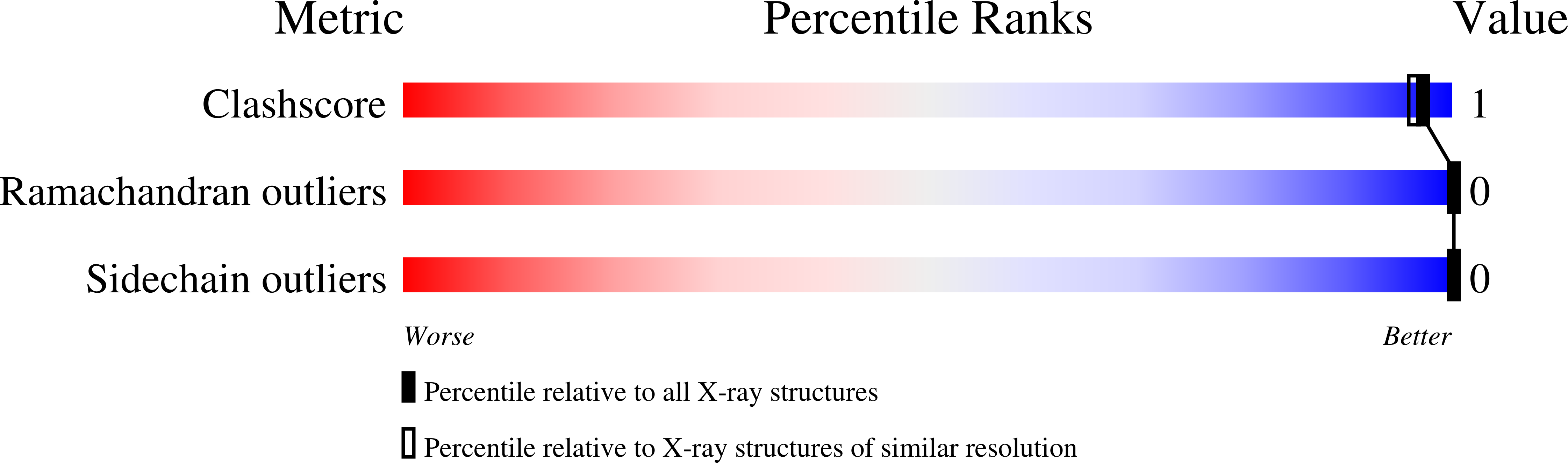
Deposition Date
2025-10-30
Release Date
2025-11-19
Last Version Date
2026-02-18
Entry Detail
PDB ID:
9YZW
Keywords:
Title:
Joint Xray/Neutron structure of Escherichia coli YajL at room temperature
Biological Source:
Source Organism(s):
Escherichia coli (Taxon ID: 562)
Expression System(s):
Method Details:
Experimental Method:
R-Value Free:
['0.16
R-Value Work:
['0.13
R-Value Observed:
['0.13
Space Group:
P 21 21 21