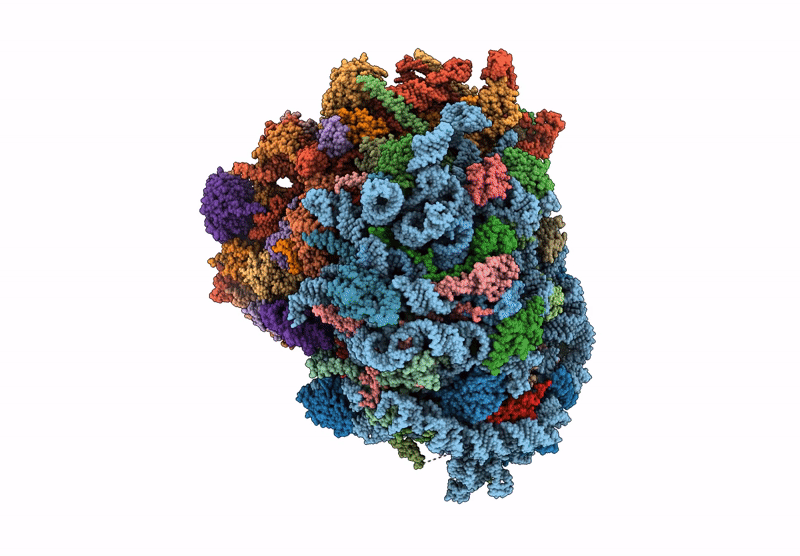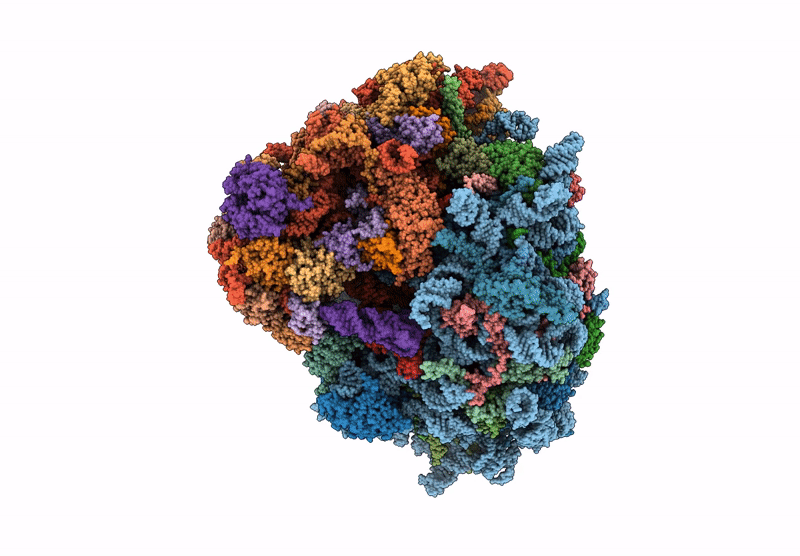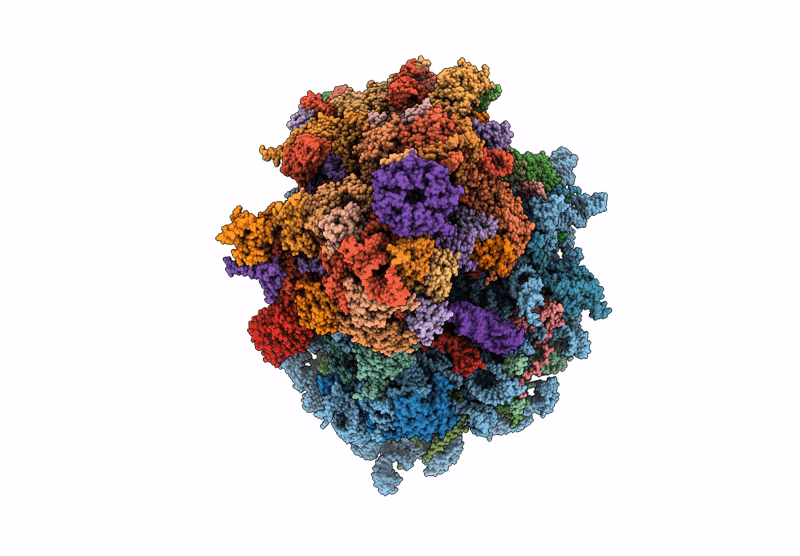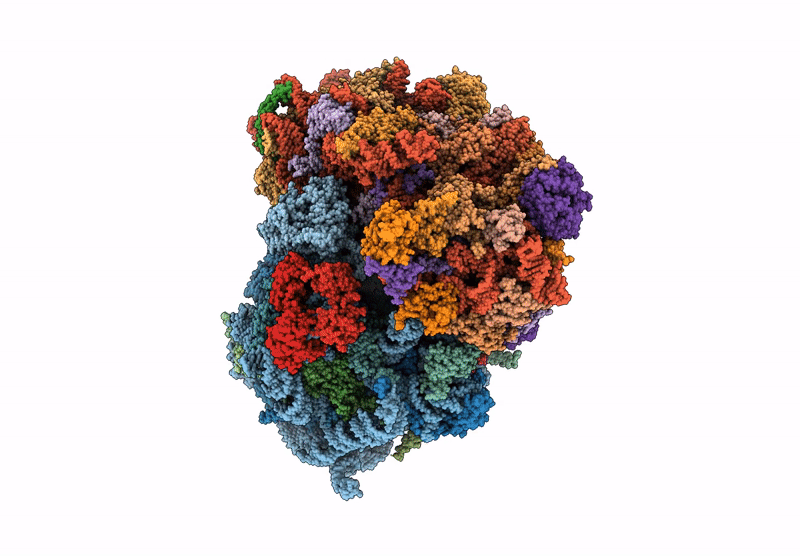
Deposition Date
2025-06-20
Release Date
2025-12-24
Last Version Date
2025-12-24
Entry Detail
PDB ID:
9P7I
Keywords:
Title:
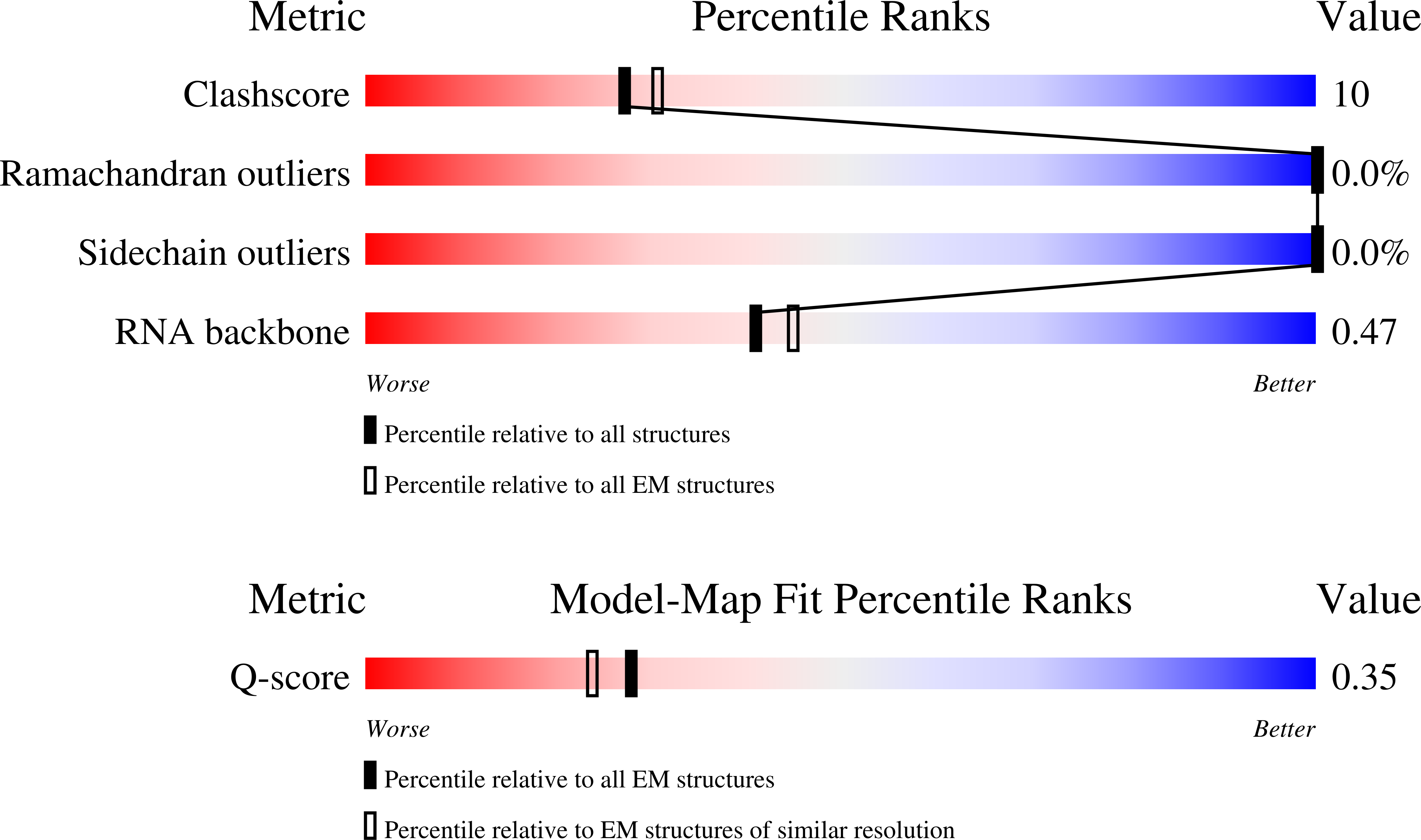
In situ HHT and CHX treated human eEF1A-A/T-P-Z state 80S ribosome
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Method Details:
Experimental Method:
Resolution:
3.69 Å
Aggregation State:
CELL
Reconstruction Method:
SINGLE PARTICLE