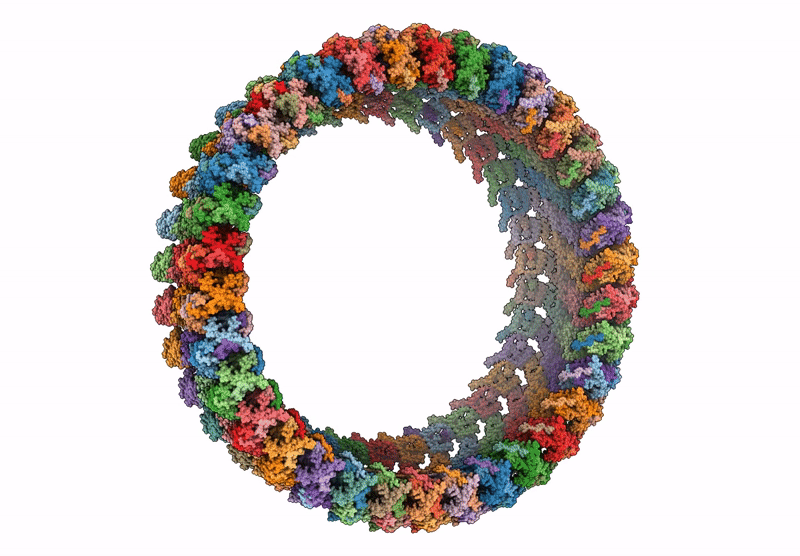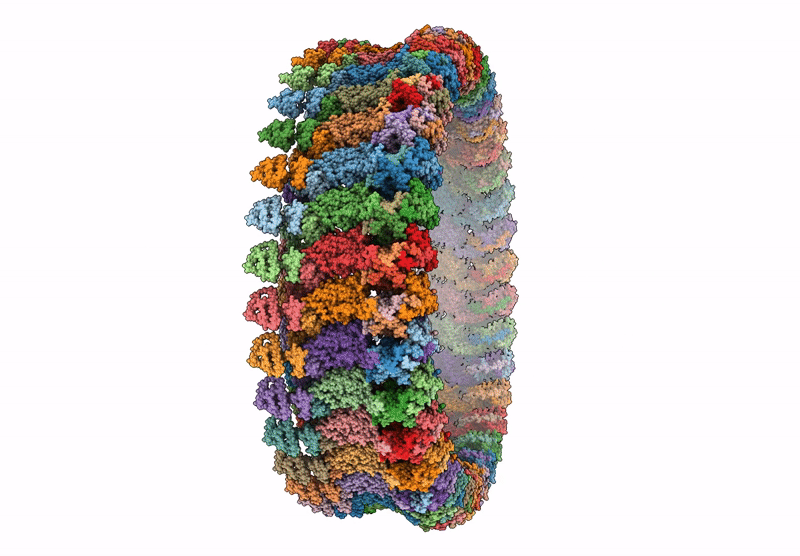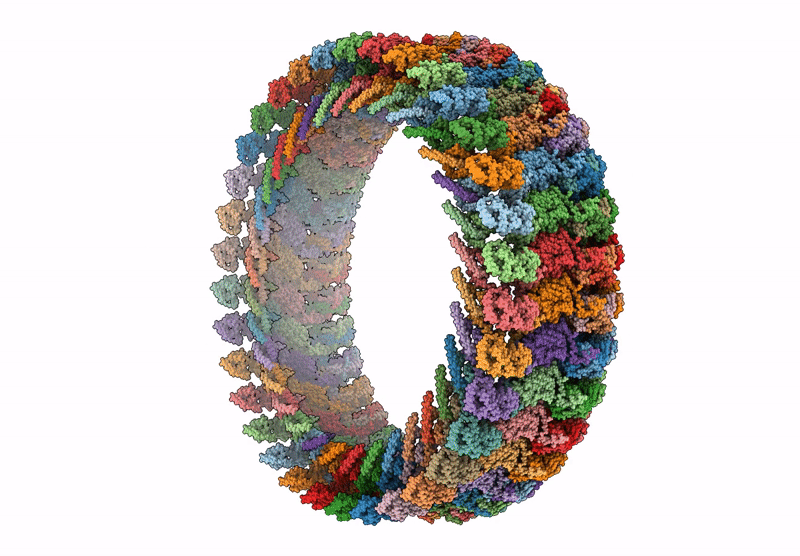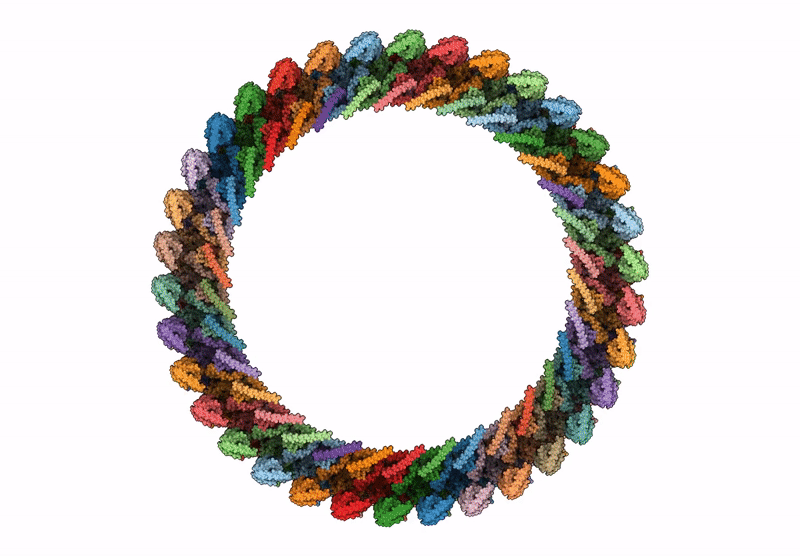
Deposition Date
2025-02-03
Release Date
2025-03-19
Last Version Date
2025-04-02
Entry Detail
PDB ID:
9N4Z
Keywords:
Title:
CCW Flagellar Switch Complex - FliF, FliG, FliM, and FliN forming 34-mer C-ring from Salmonella
Biological Source:
Source Organism(s):
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.00 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE