Deposition Date
2024-05-07
Release Date
2024-10-09
Last Version Date
2025-01-01
Entry Detail
PDB ID:
9F93
Keywords:
Title:
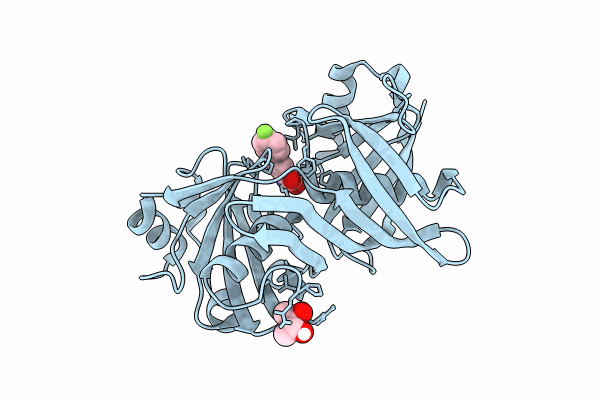
Complex of phenazine biosynthesis enzyme PhzF with 2-amino-5-(4-fluorophenyl)benzoic acid
Biological Source:
Source Organism(s):
Pseudomonas fluorescens (Taxon ID: 294)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.54 Å
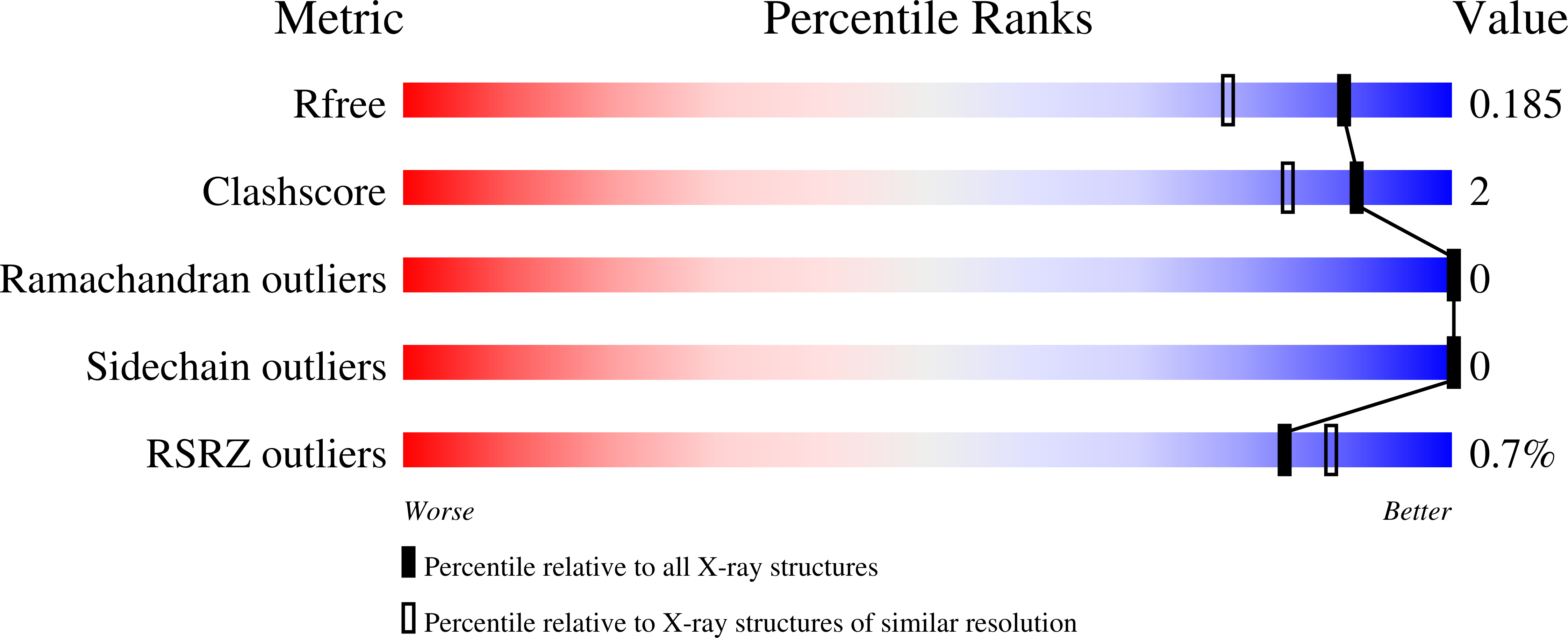
R-Value Free:
0.18
R-Value Work:
0.16
R-Value Observed:
0.16
Space Group:
P 32 2 1


