Deposition Date
2024-08-01
Release Date
2025-08-06
Last Version Date
2025-12-03
Entry Detail
PDB ID:
9CY7
Keywords:
Title:
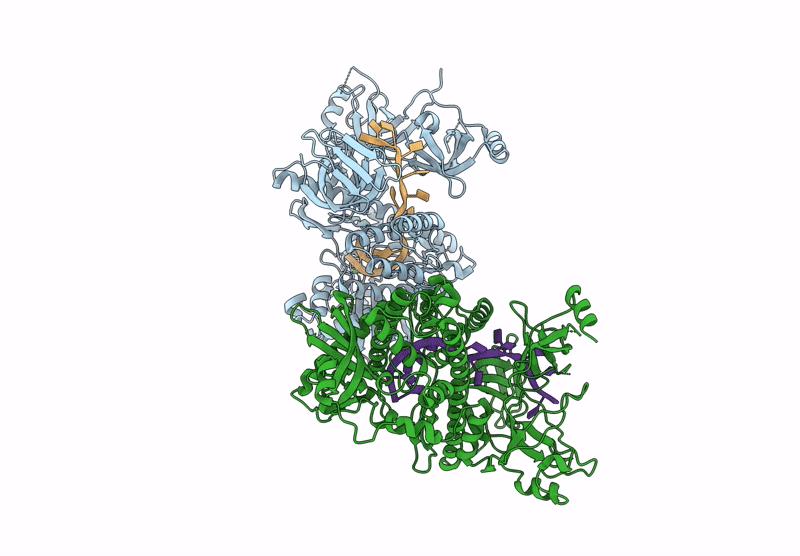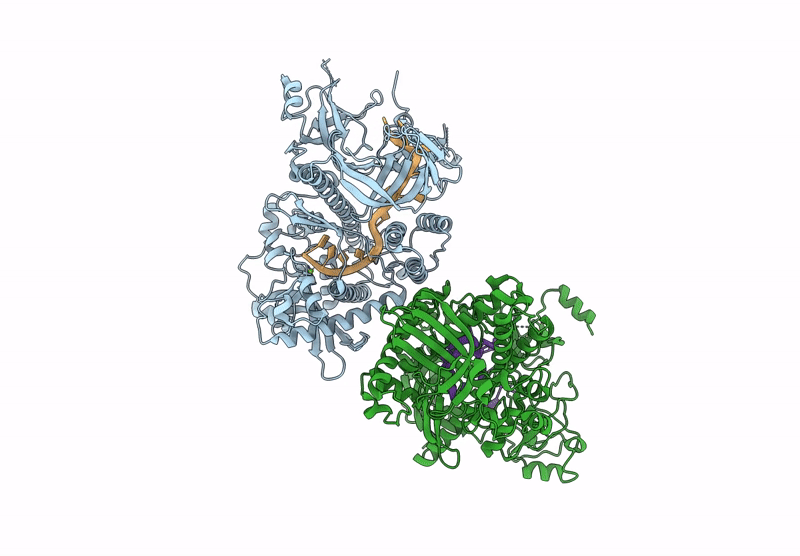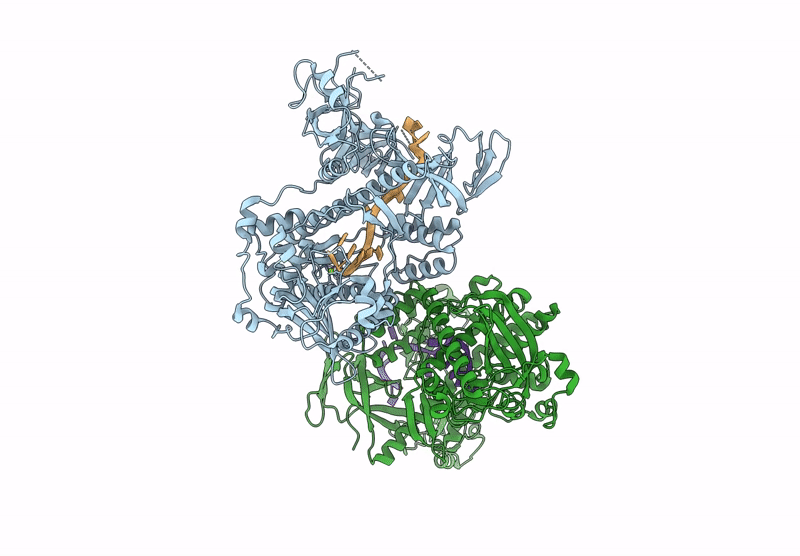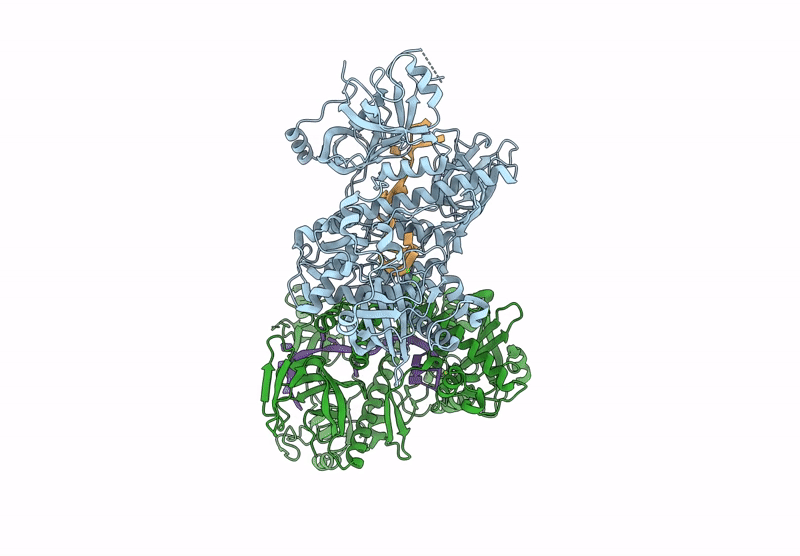
Structure of S.pombe Dis3L2 in complex with oligoU RNA substrate
Biological Source:
Source Organism(s):
Schizosaccharomyces pombe (strain 972 / ATCC 24843) (Taxon ID: 284812)
Schizosaccharomyces pombe 927 (Taxon ID: 1264690)
Schizosaccharomyces pombe 927 (Taxon ID: 1264690)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.52 Å
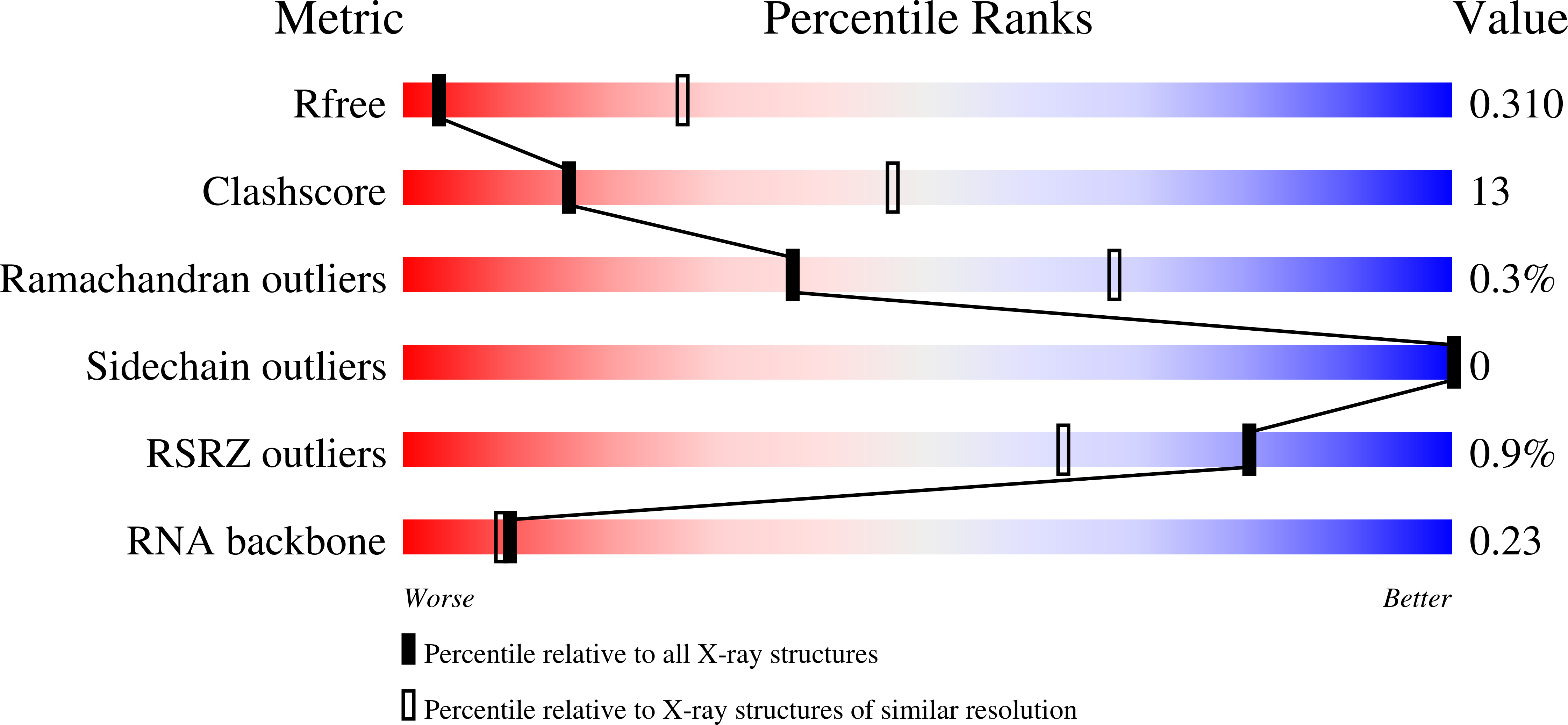
R-Value Free:
0.31
R-Value Work:
0.28
R-Value Observed:
0.28
Space Group:
P 41 21 2