Deposition Date
2024-03-21
Release Date
2025-08-27
Last Version Date
2025-09-10
Entry Detail
PDB ID:
9B50
Keywords:
Title:
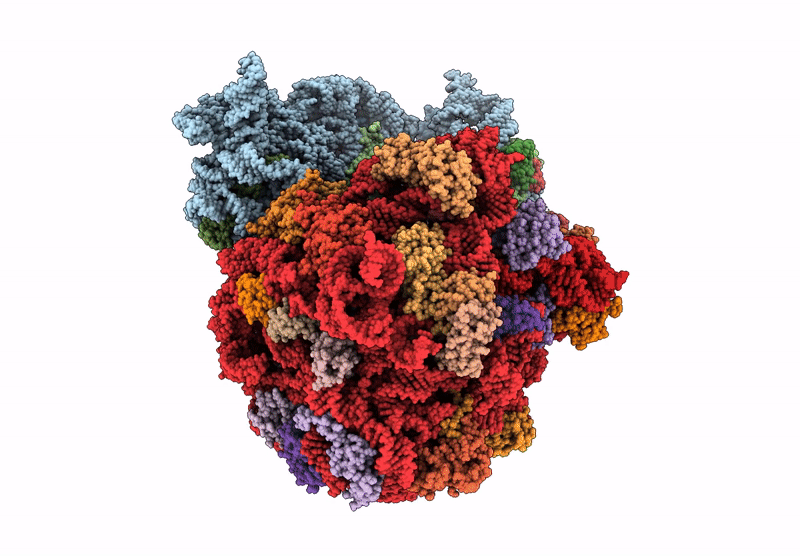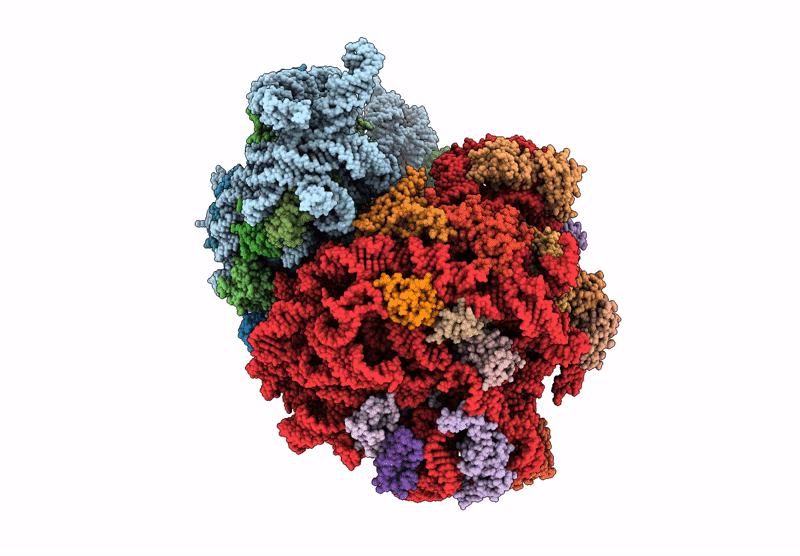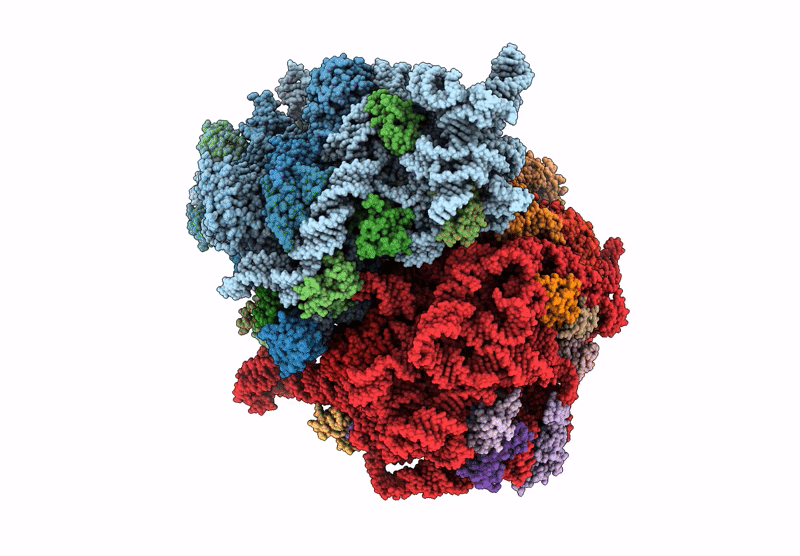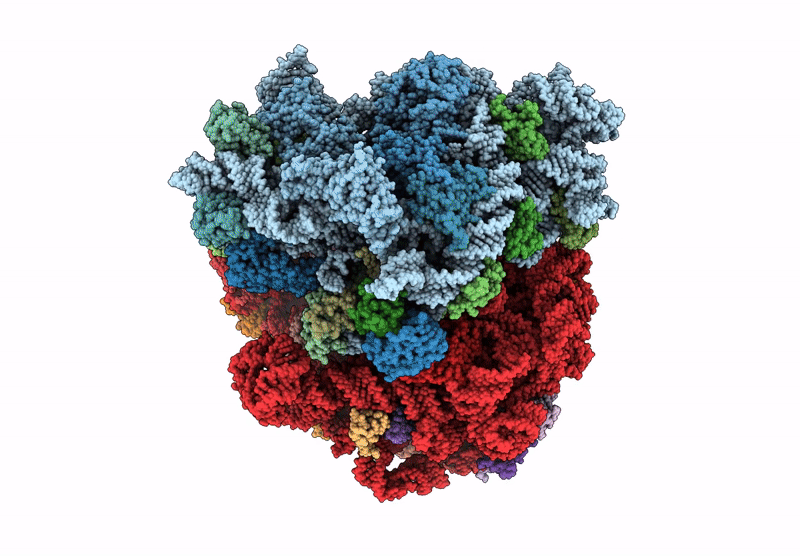
E. coli 70S ribosome complex (unmethylated 16S A1408 + arbekacin)
Biological Source:
Source Organism(s):
Escherichia coli (Taxon ID: 562)
Method Details:
Experimental Method:
Resolution:
2.70 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE