Deposition Date
2024-06-17
Release Date
2024-09-04
Last Version Date
2025-03-19
Entry Detail
PDB ID:
8ZYH
Keywords:
Title:
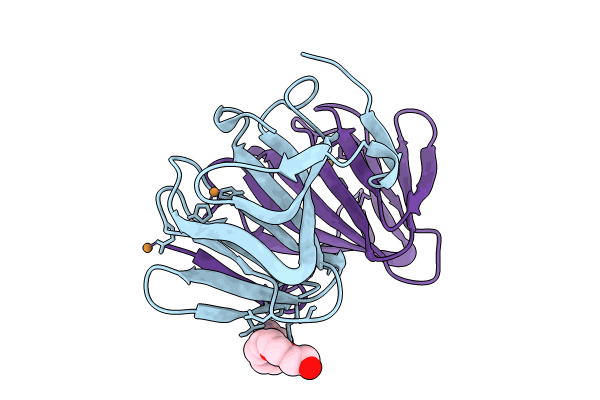
Crystal structure of a cupin protein (tm1459, I49C-4py/H52A/H54A/C106D mutant) in copper (Cu) substituted form
Biological Source:
Source Organism(s):
Thermotoga maritima MSB8 (Taxon ID: 243274)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.07 Å
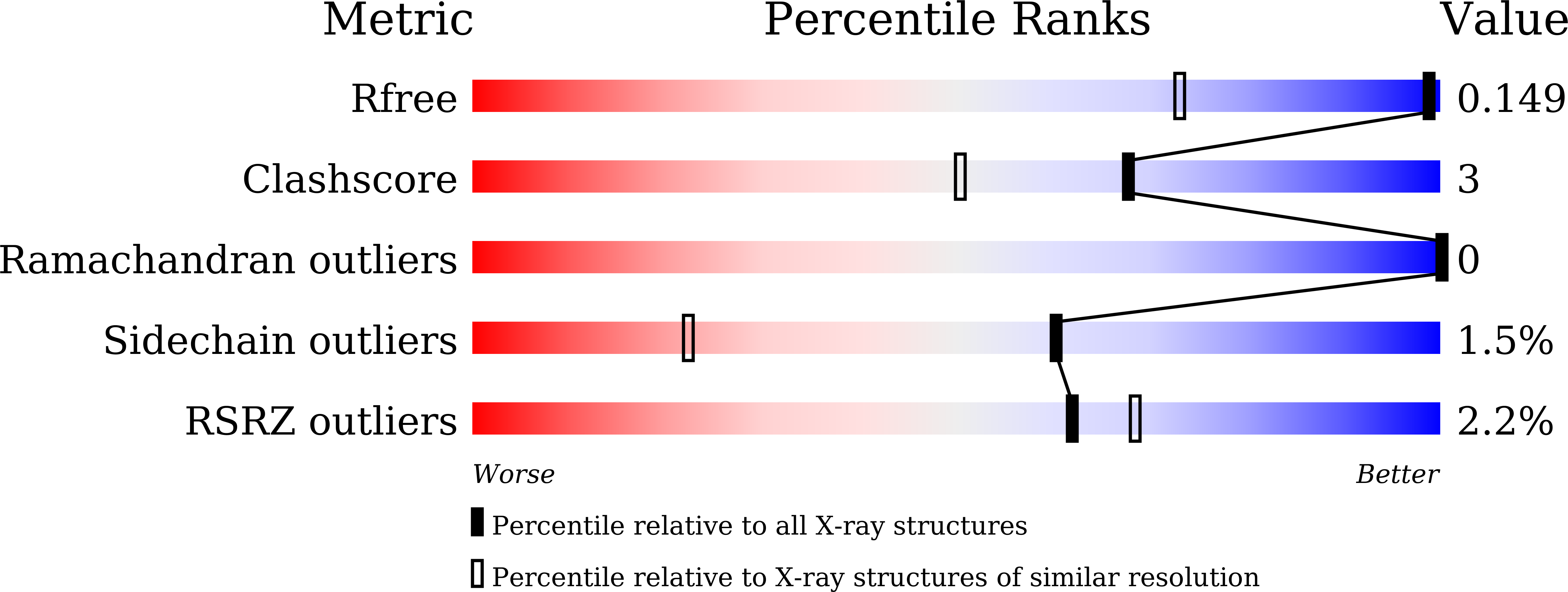
R-Value Free:
0.16
R-Value Work:
0.13
R-Value Observed:
0.13
Space Group:
P 21 21 21