Deposition Date
2023-05-25
Release Date
2023-09-13
Last Version Date
2023-10-18
Entry Detail
PDB ID:
8SYD
Keywords:
Title:
X-ray crystal structure of UDP-2,3-diacetamido-2,3-dideoxy-glucuronic acid-2-epimerase from Thermus thermophilus strain HB27, D98N variant in the presence of UDP-2,3-diacetamido-2,3-dideoxy-glucuronic acid and UDP-N-acetylglucosamine at pH 6
Biological Source:
Source Organism(s):
Thermus thermophilus HB27 (Taxon ID: 262724)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.20 Å
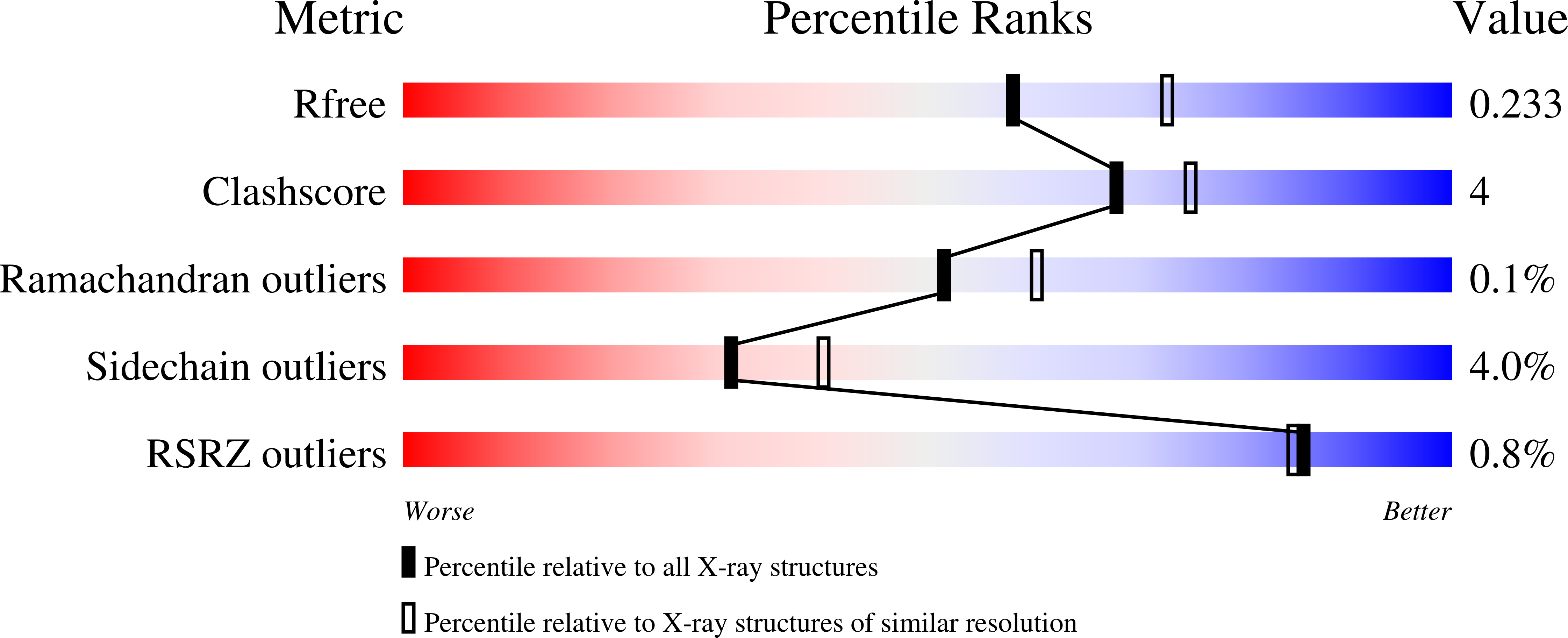
R-Value Free:
0.22
R-Value Work:
0.17
R-Value Observed:
0.17
Space Group:
P 1 21 1


