Deposition Date
2023-03-13
Release Date
2023-10-18
Last Version Date
2023-12-27
Entry Detail
PDB ID:
8OF0
Keywords:
Title:
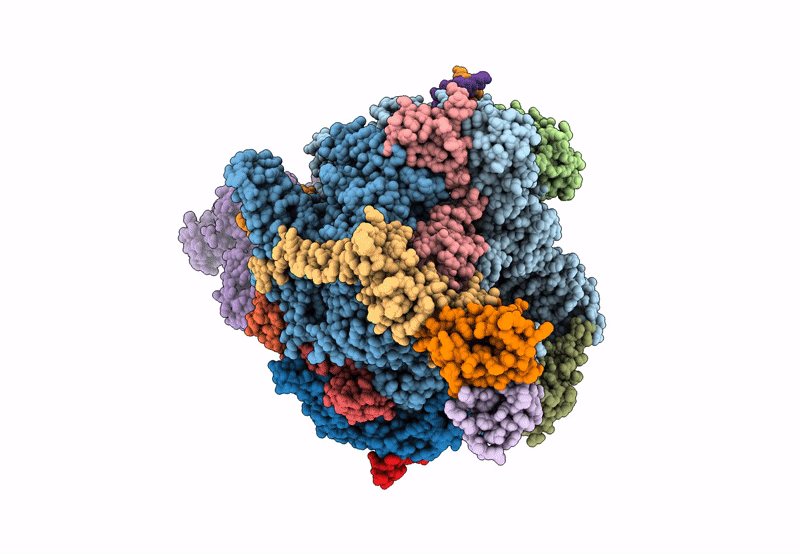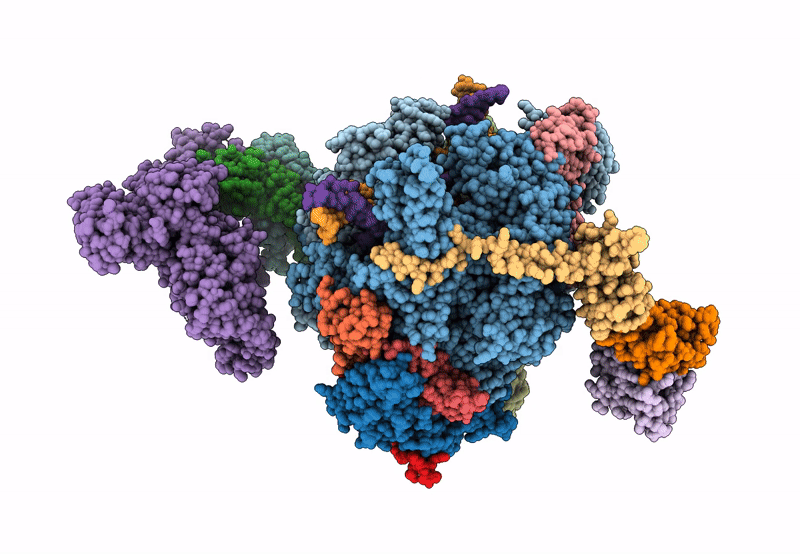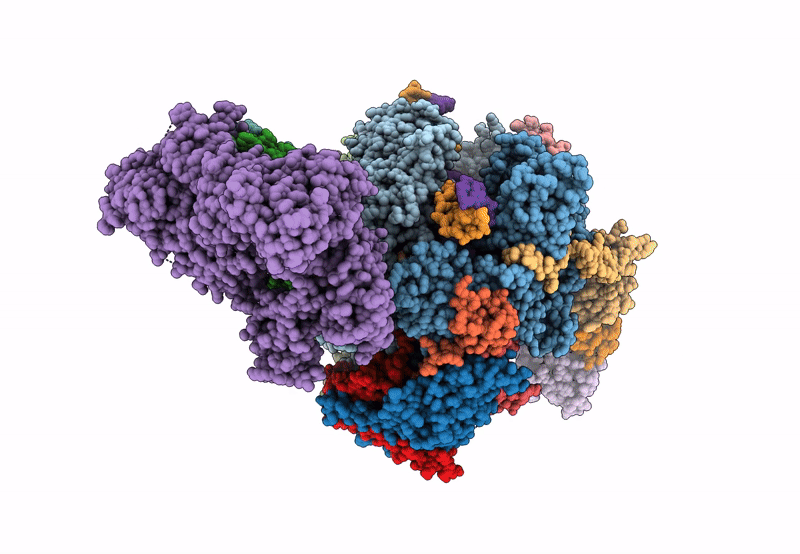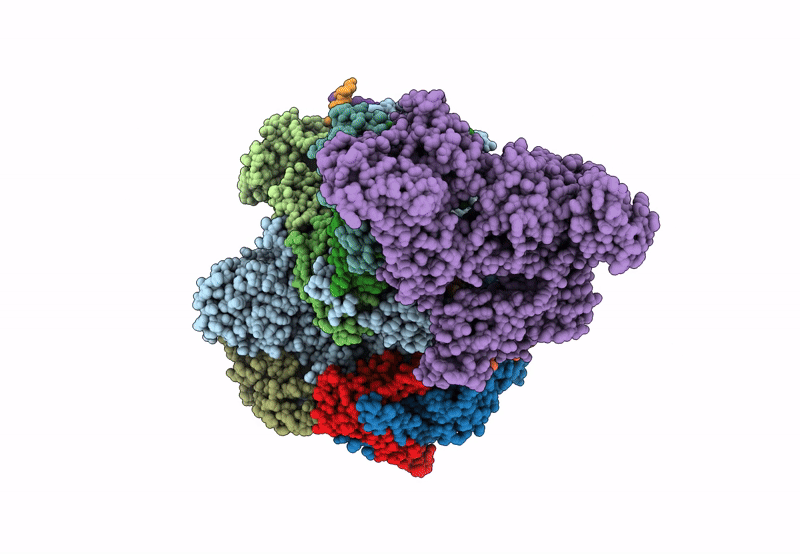
Structure of the mammalian Pol II-SPT6-Elongin complex, Structure 1
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
synthetic construct (Taxon ID: 32630)
Sus scrofa (Taxon ID: 9823)
synthetic construct (Taxon ID: 32630)
Sus scrofa (Taxon ID: 9823)
Expression System(s):
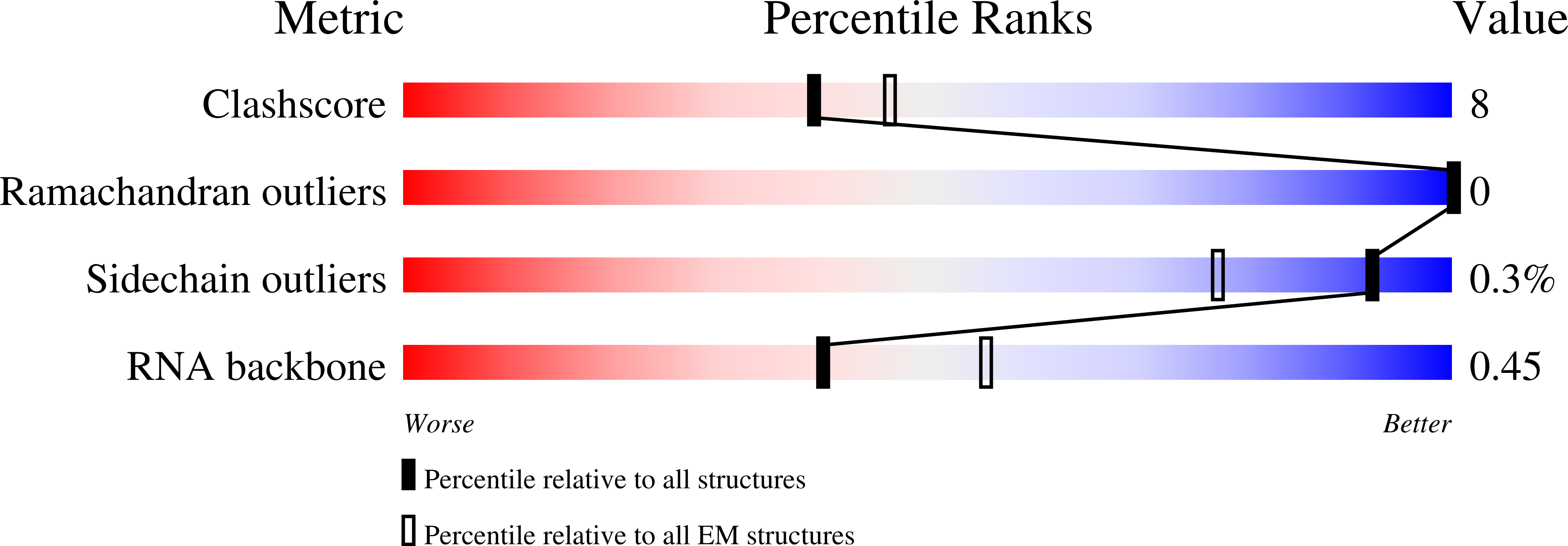
Method Details:
Experimental Method:
Resolution:
3.05 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE