Deposition Date
2023-04-21
Release Date
2023-07-26
Last Version Date
2023-11-01
Entry Detail
PDB ID:
8J52
Keywords:
Title:
Crystal structure of Flavihumibacter petaseus GH31 alpha-galactosidase mutant D304A in complex with alpha-1,4-galactobiose
Biological Source:
Source Organism(s):
Flavihumibacter petaseus NBRC 106054 (Taxon ID: 1220578)
Expression System(s):
Method Details:
Experimental Method:
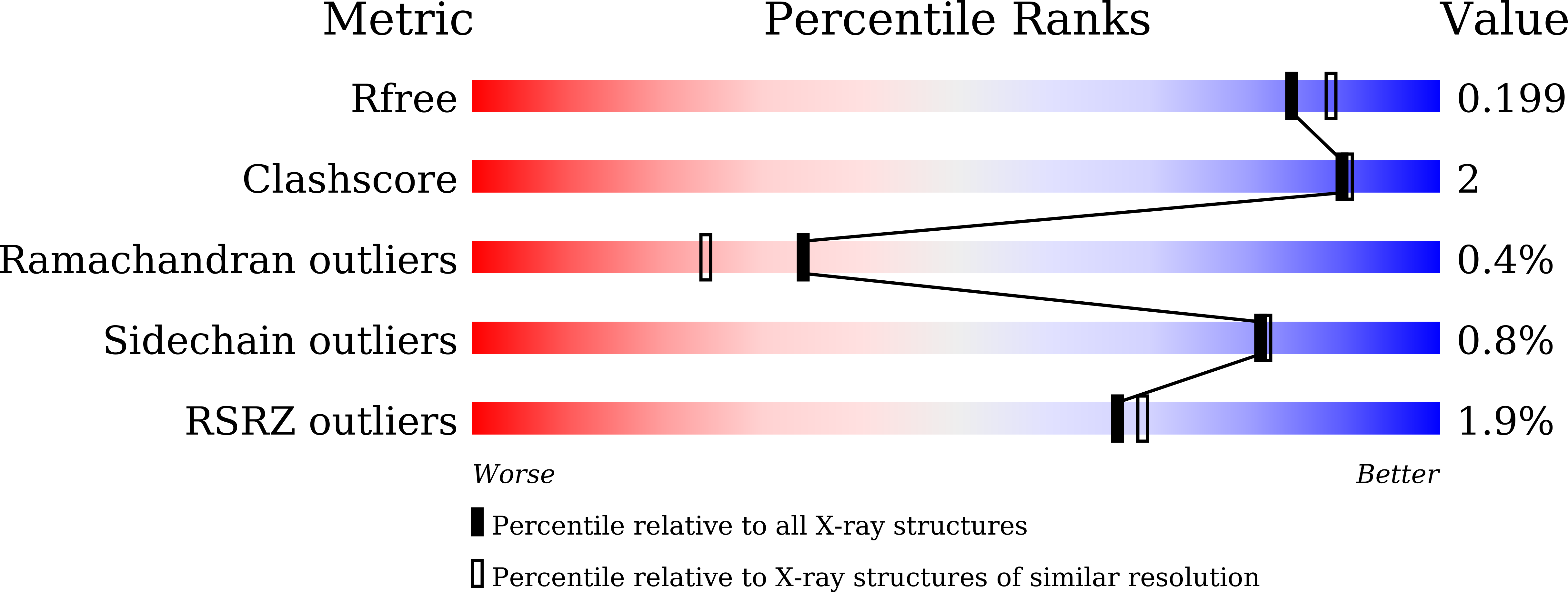
Resolution:
1.90 Å
R-Value Free:
0.19
R-Value Work:
0.13
Space Group:
P 1


