Deposition Date
1990-10-26
Release Date
1993-10-31
Last Version Date
2023-11-15
Entry Detail
PDB ID:
8HVP
Keywords:
Title:
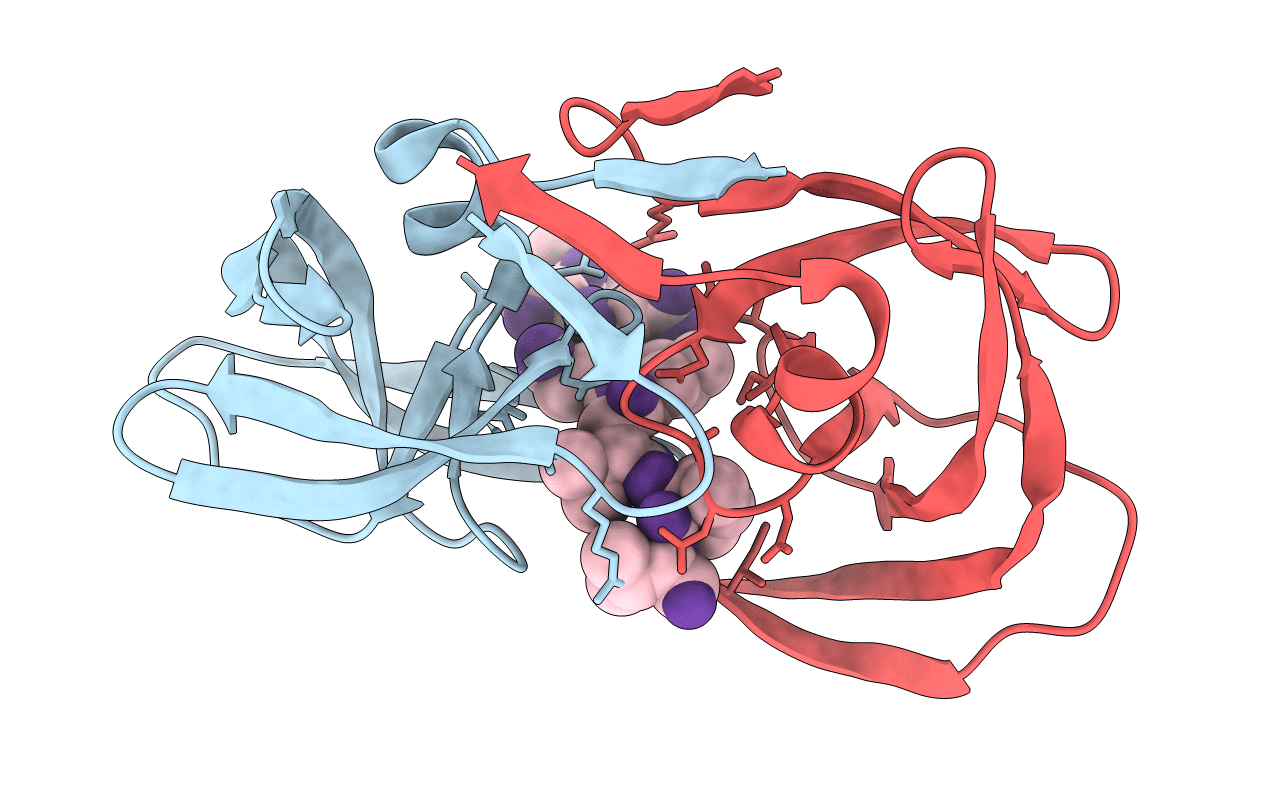
STRUCTURE AT 2.5-ANGSTROMS RESOLUTION OF CHEMICALLY SYNTHESIZED HUMAN IMMUNODEFICIENCY VIRUS TYPE 1 PROTEASE COMPLEXED WITH A HYDROXYETHYLENE*-BASED INHIBITOR
Biological Source:
Source Organism(s):
Human immunodeficiency virus 1 (Taxon ID: 11676)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.50 Å
R-Value Observed:
0.13
Space Group:
P 21 21 21


