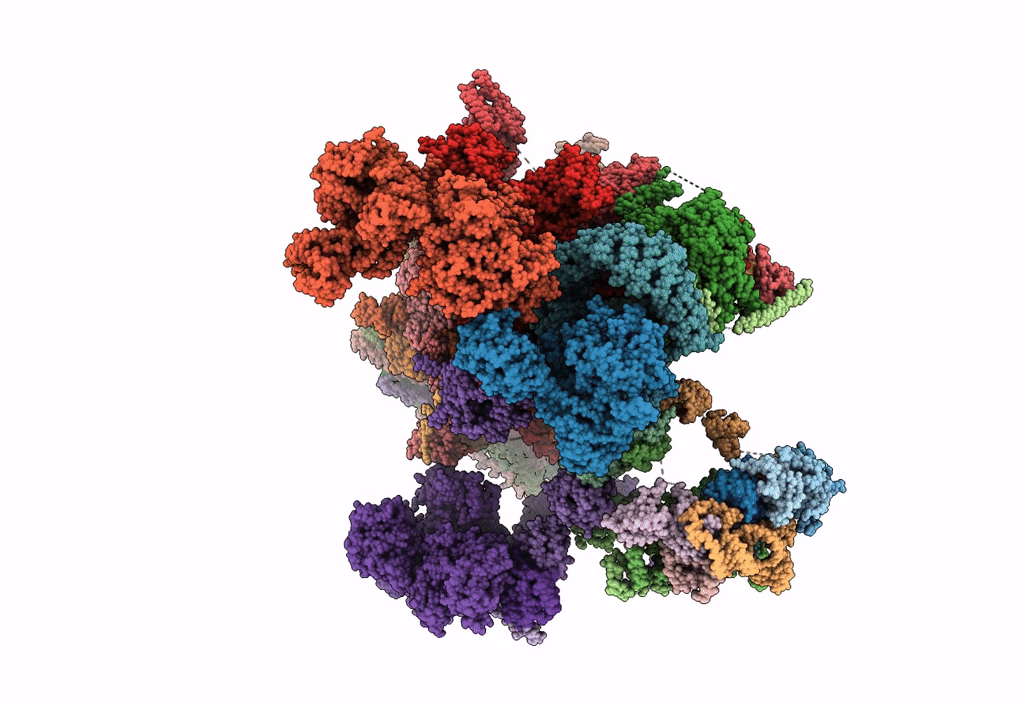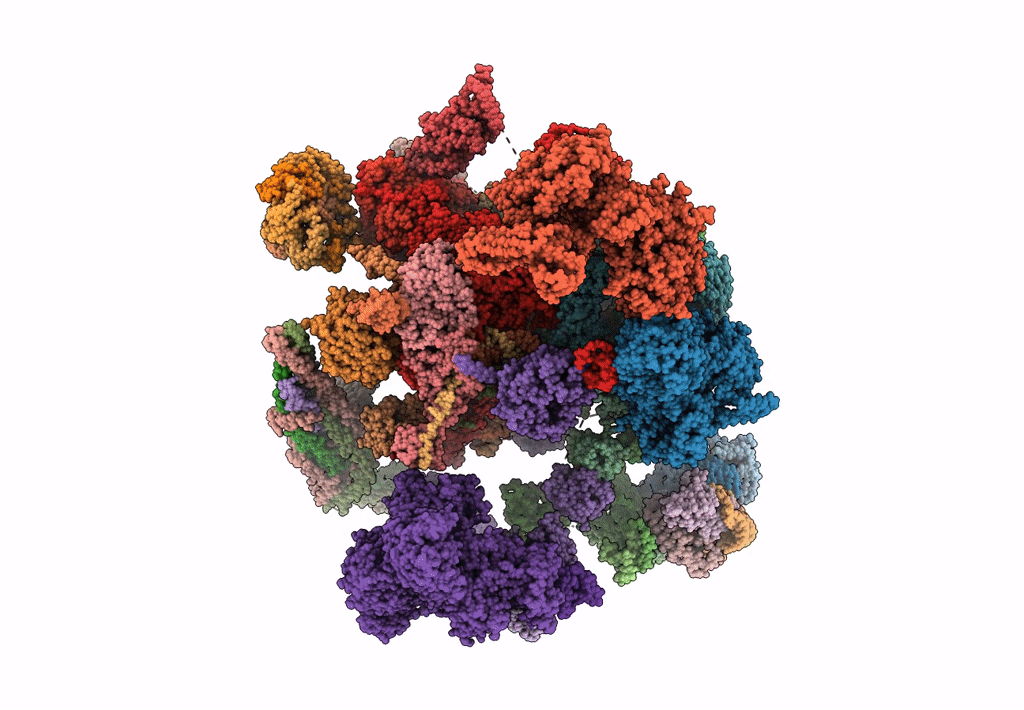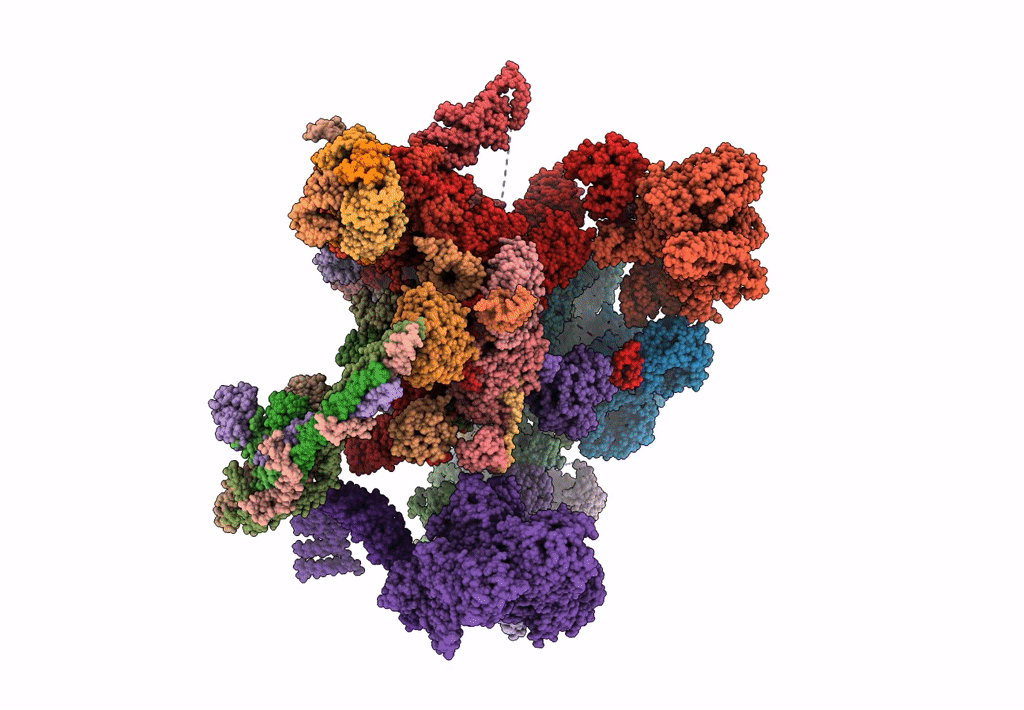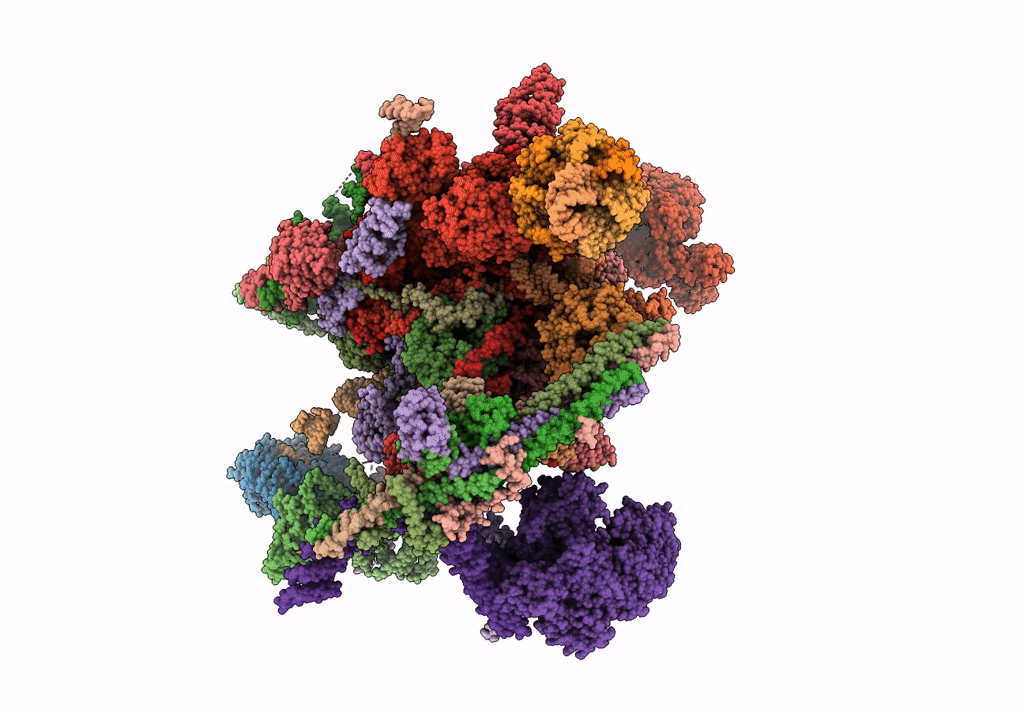
Deposition Date
2023-02-07
Release Date
2023-05-10
Last Version Date
2024-07-24
Entry Detail
PDB ID:
8CH6
Keywords:
Title:
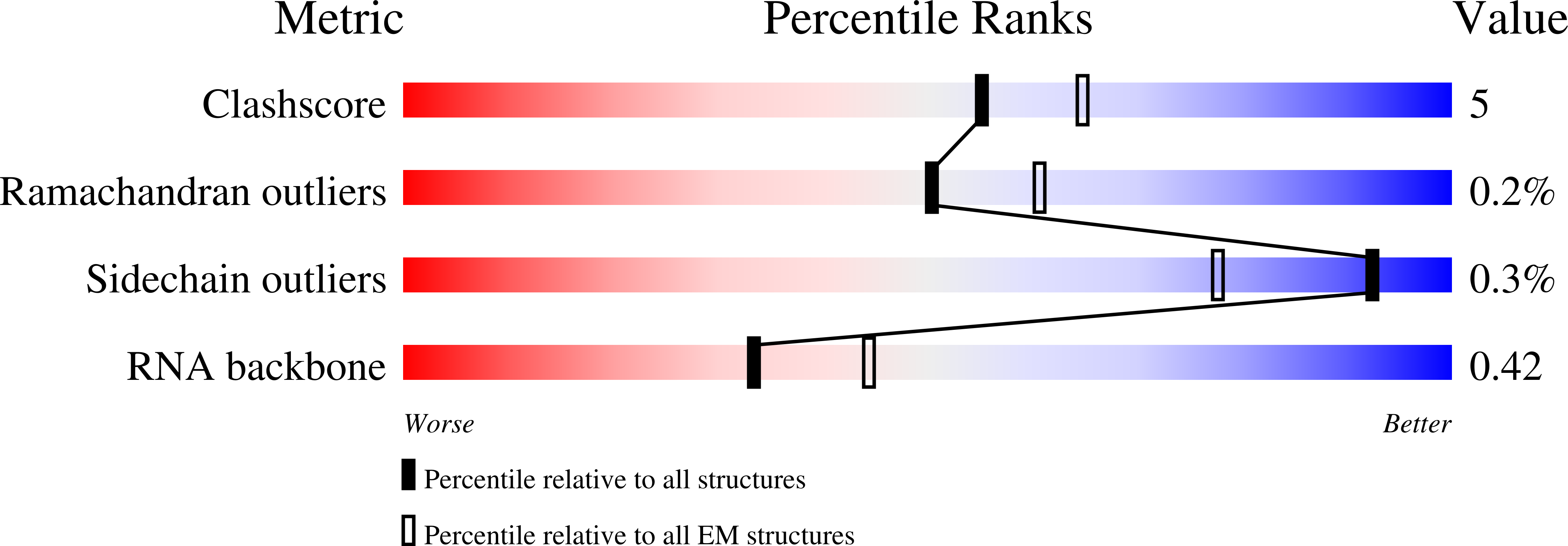
Structure of a late-stage activated spliceosome (BAqr) arrested with a dominant-negative Aquarius mutant (state B complex).
Biological Source:
Source Organism(s):
unidentified adenovirus (Taxon ID: 10535)
Homo sapiens (Taxon ID: 9606)
Homo sapiens (Taxon ID: 9606)
Method Details:
Experimental Method:
Resolution:
5.90 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE