Deposition Date
2022-04-12
Release Date
2023-04-26
Last Version Date
2024-11-06
Entry Detail
PDB ID:
7ZKD
Keywords:
Title:
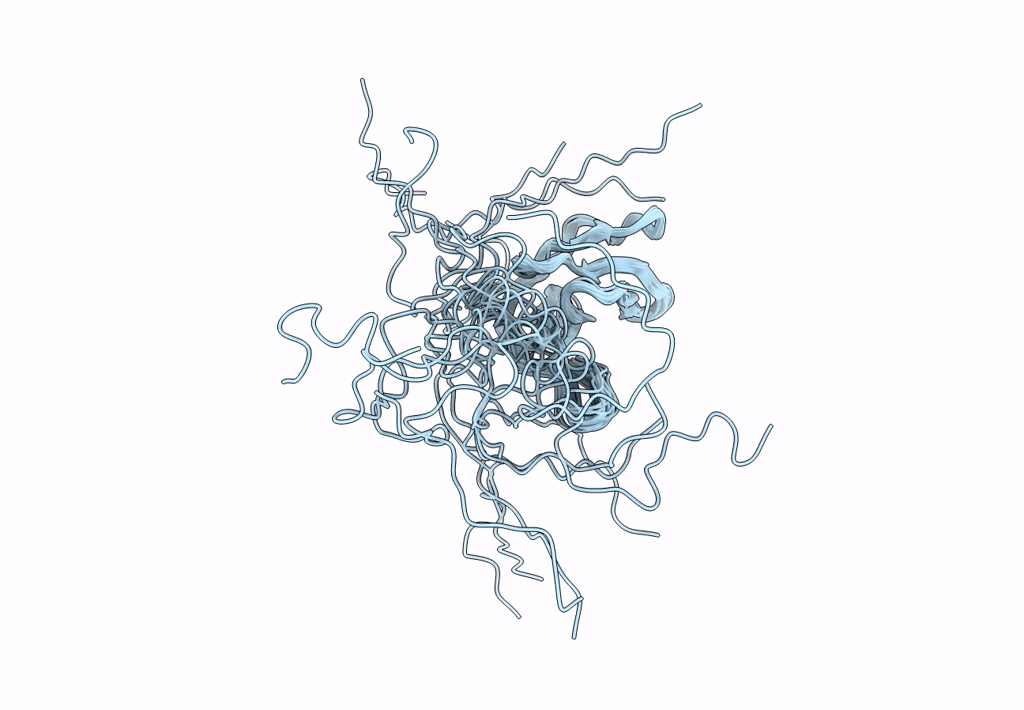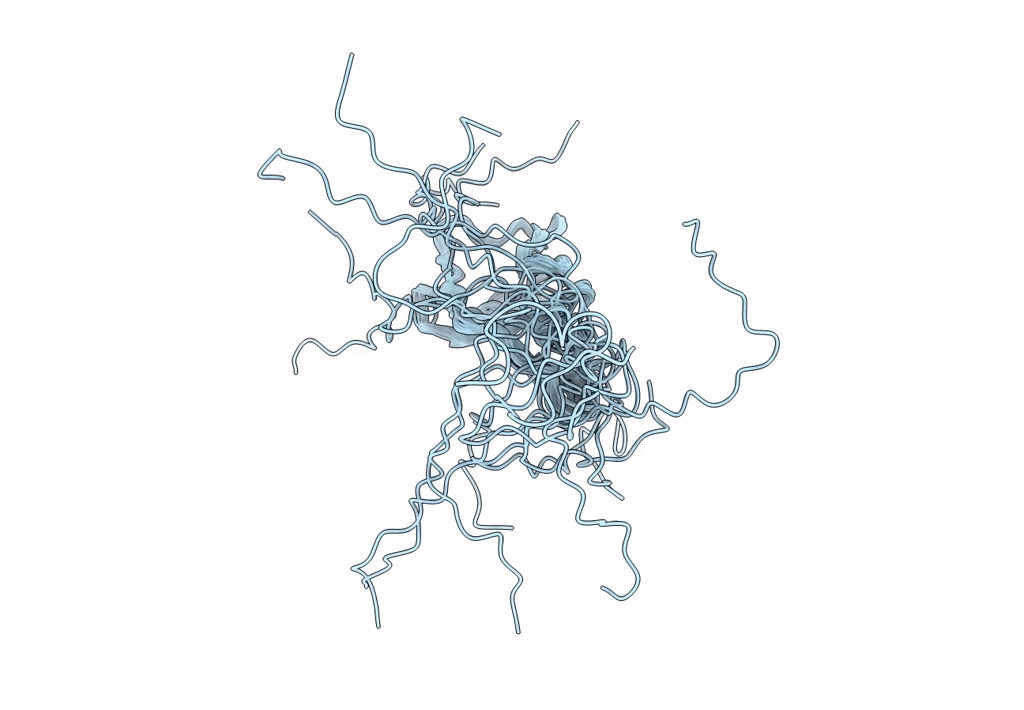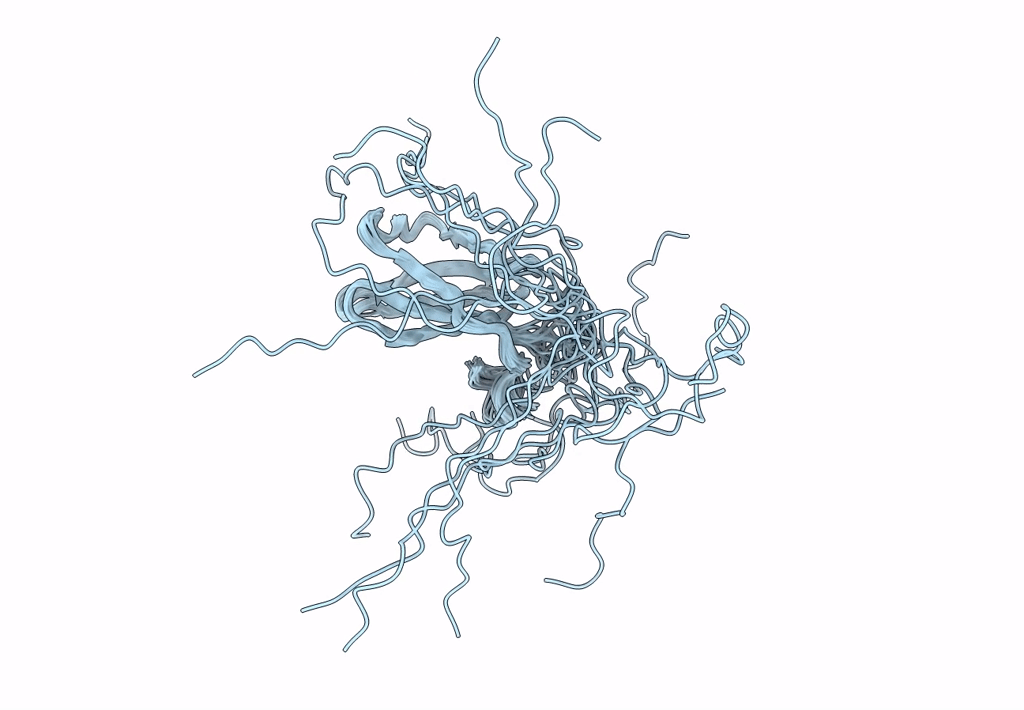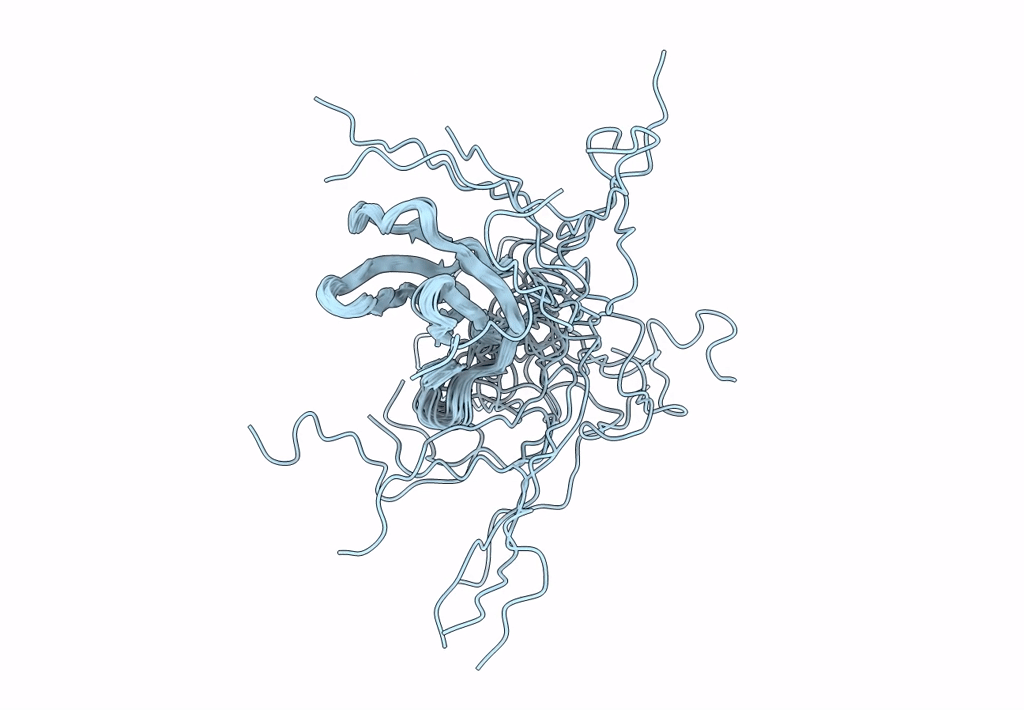
The NMR structure of the MAX47 effector from Magnaporthe Oryzae
Biological Source:
Source Organism(s):
Pyricularia oryzae (Taxon ID: 318829)
Expression System(s):
Method Details:
Experimental Method:
Conformers Calculated:
20
Conformers Submitted:
20
Selection Criteria:
structures with the lowest energy