Deposition Date
2021-12-10
Release Date
2022-01-12
Last Version Date
2024-02-28
Entry Detail
PDB ID:
7T4M
Keywords:
Title:
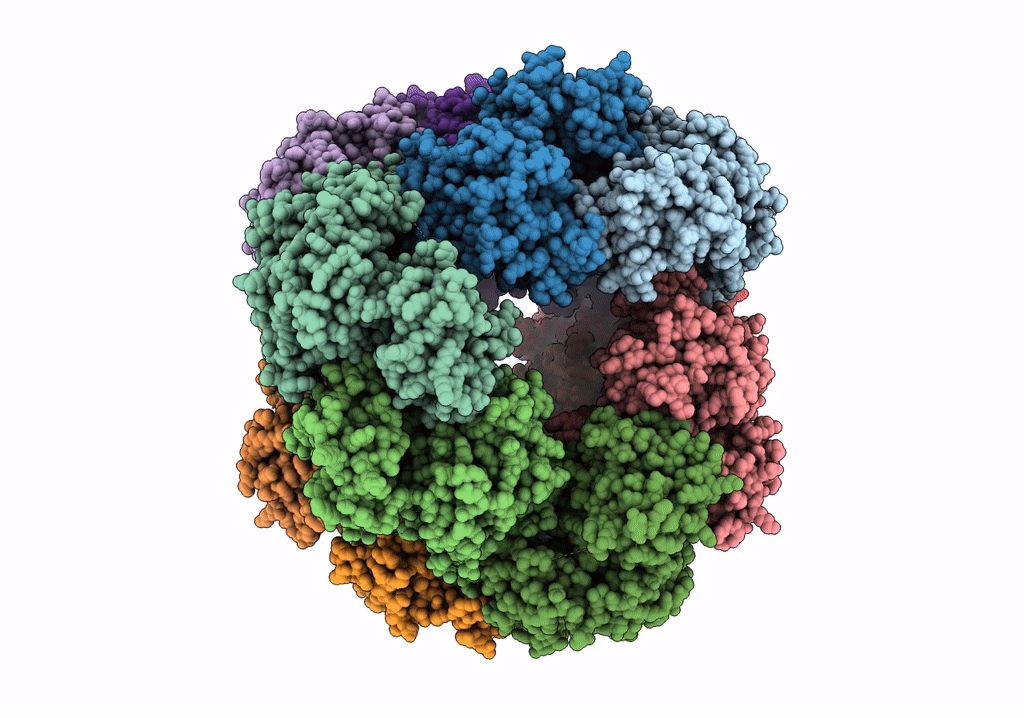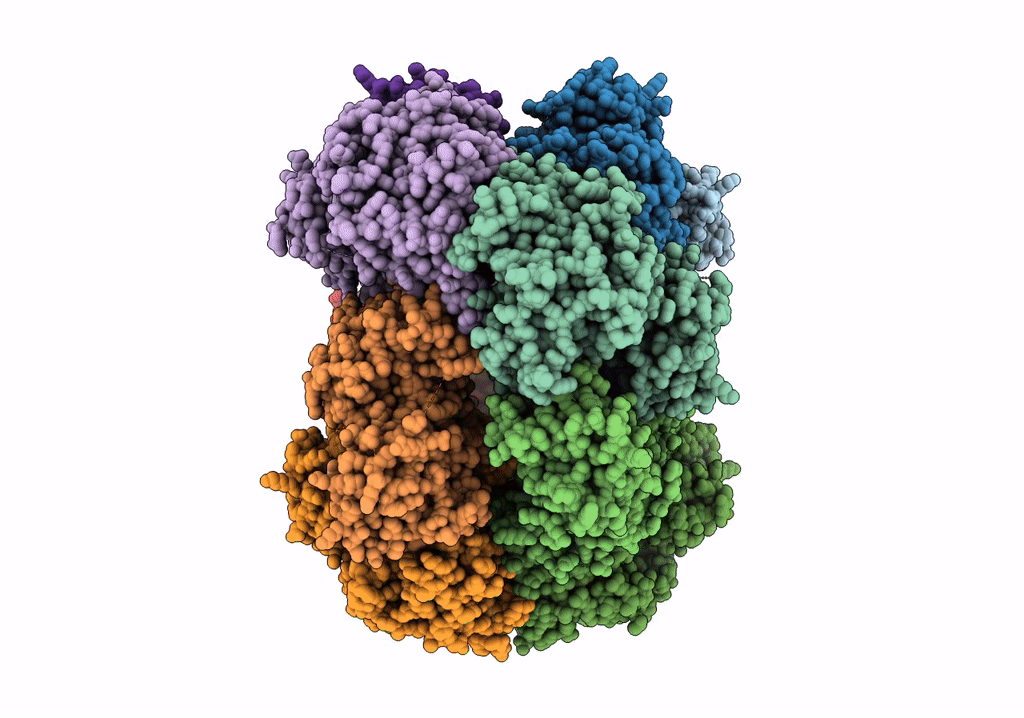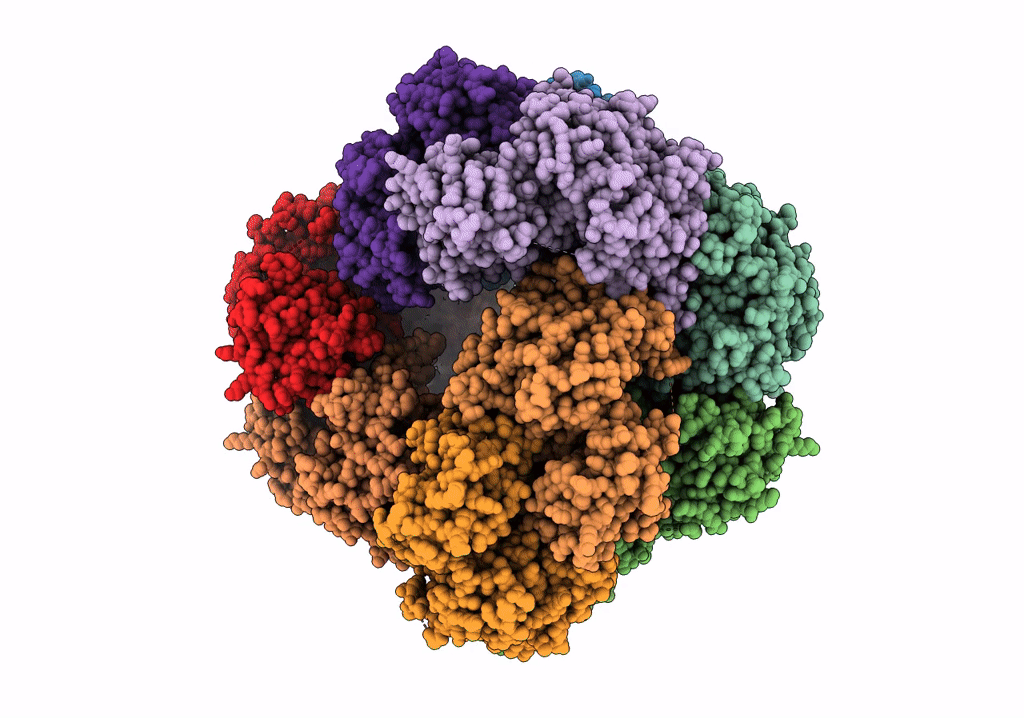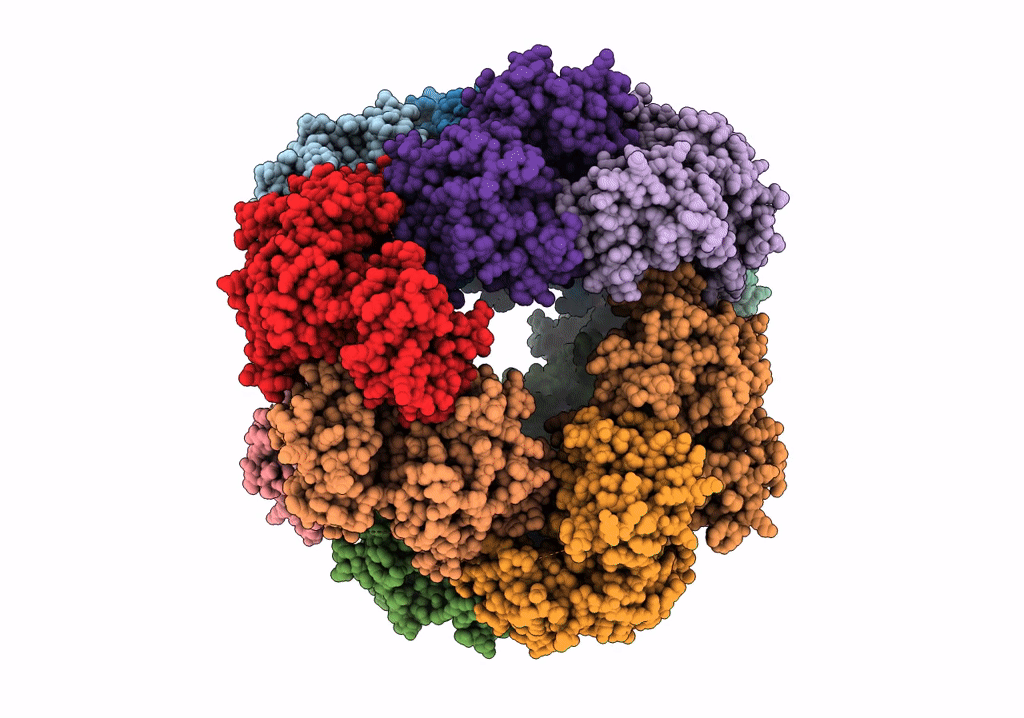
Structure of dodecameric unphosphorylated Pediculus humanus (Ph) PINK1 D357A mutant
Biological Source:
Source Organism(s):
Pediculus humanus corporis (Taxon ID: 121224)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.48 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE