Deposition Date
2020-06-29
Release Date
2020-09-30
Last Version Date
2024-01-31
Entry Detail
PDB ID:
6ZJK
Keywords:
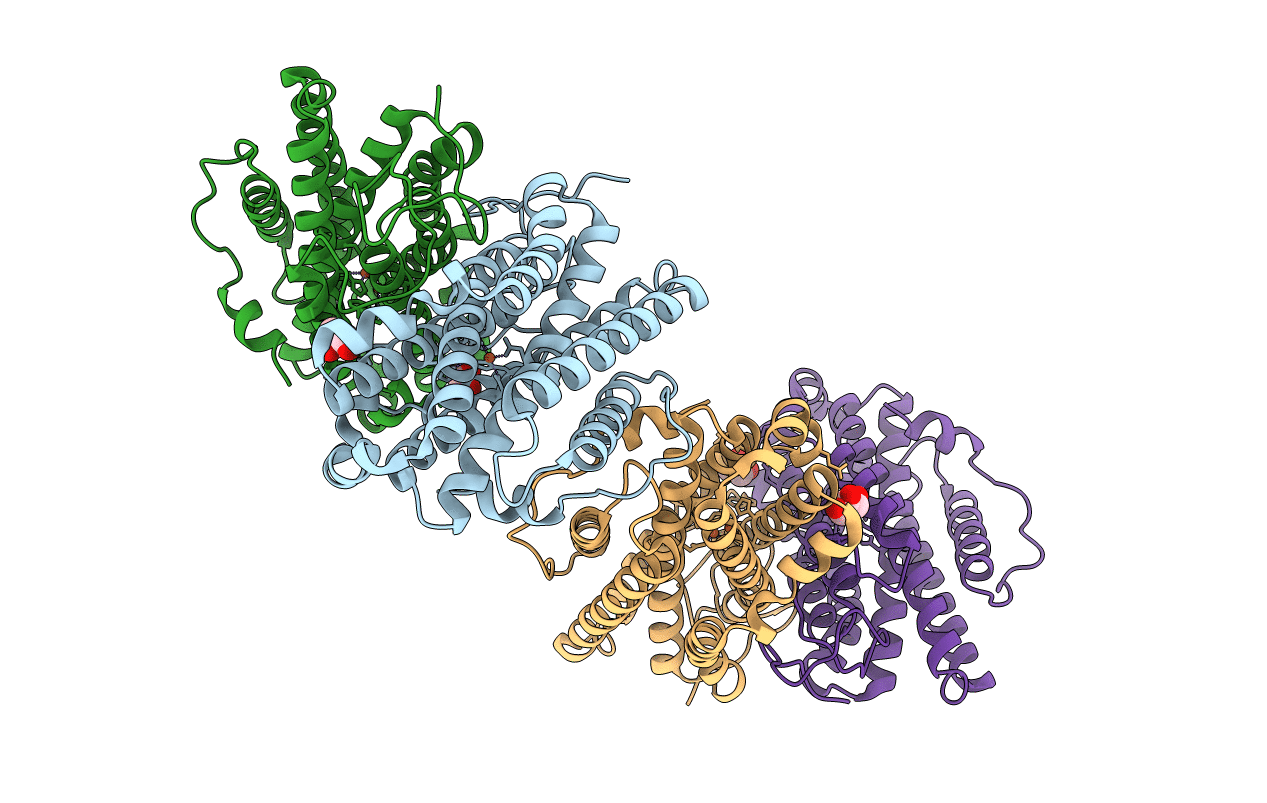
Title:
Ribonucleotide reductase R2 subunit from Clostridium botulinum
Biological Source:
Source Organism(s):
Expression System(s):
Method Details:
Experimental Method:
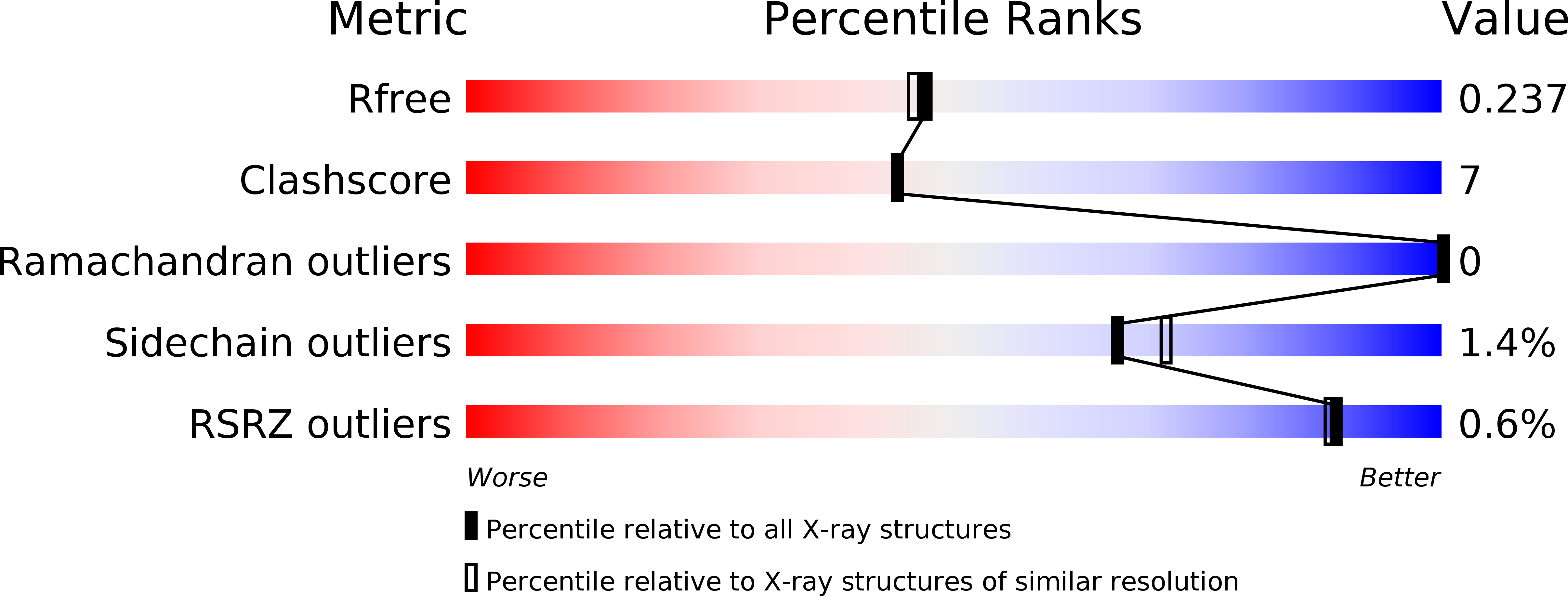
Resolution:
2.00 Å
R-Value Free:
0.23
R-Value Work:
0.22
R-Value Observed:
0.22
Space Group:
C 1 2 1