Deposition Date
2020-06-03
Release Date
2020-11-04
Last Version Date
2024-10-09
Entry Detail
PDB ID:
6X9U
Keywords:
Title:
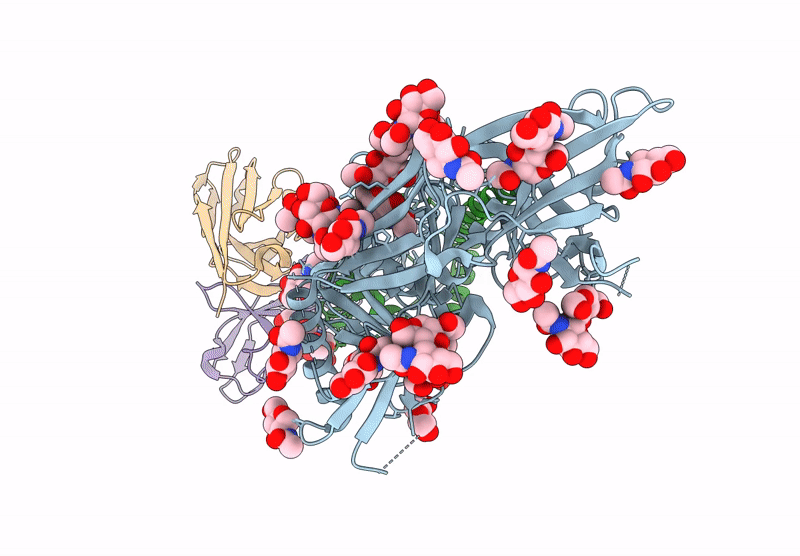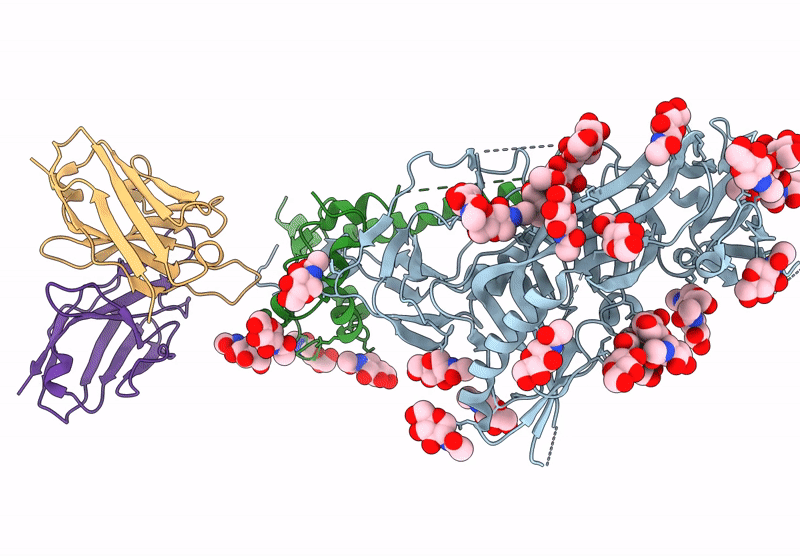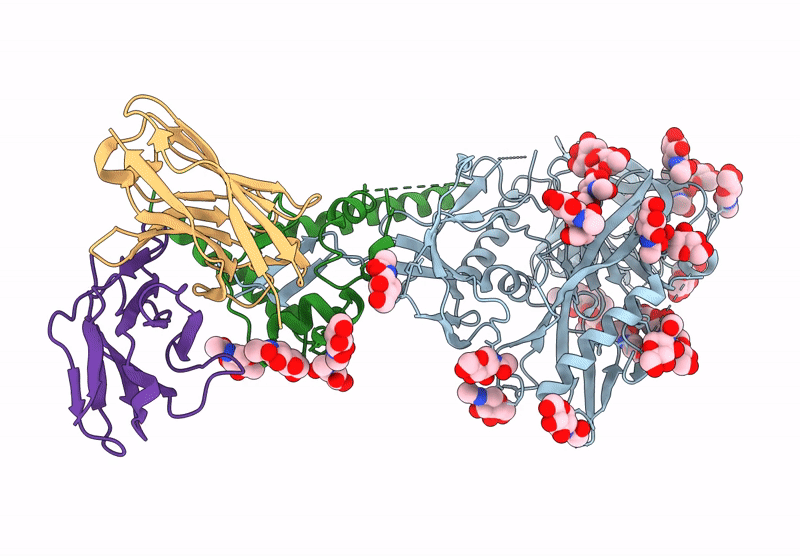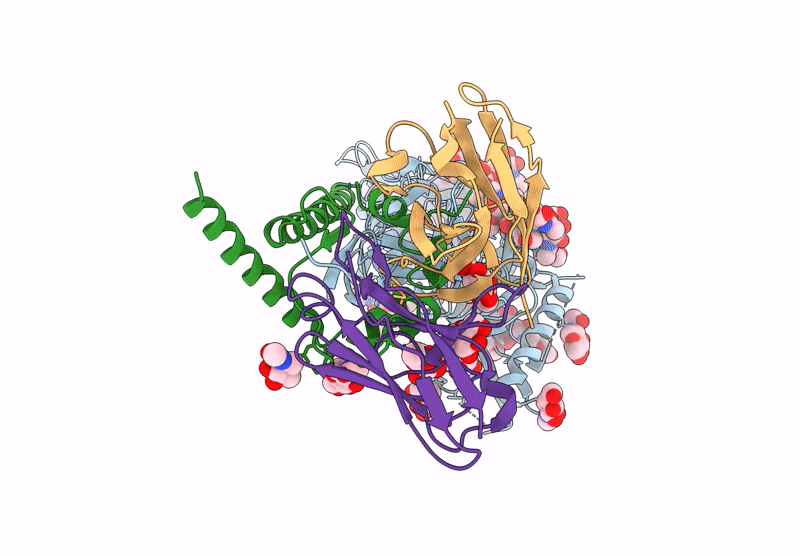
HIV-1 Envelope Glycoprotein BG505 SOSIP.664, expressed in HEK293S cells and partially deglycosylated by endoglycosidase H, in complex with RM20A3 Fab
Biological Source:
Source Organism:
Human immunodeficiency virus 1 (Taxon ID: 11676)
Macaca mulatta (Taxon ID: 9544)
Macaca mulatta (Taxon ID: 9544)
Host Organism:
Method Details:
Experimental Method:
Resolution:
3.20 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE