Deposition Date
2020-05-13
Release Date
2020-09-02
Last Version Date
2023-10-18
Entry Detail
PDB ID:
6WYO
Keywords:
Title:
Crystal structure of Danio rerio histone deacetylase 6 catalytic domain 1 (CD1) H82F F202Y double mutant complexed with Trichostatin A
Biological Source:
Source Organism:
Danio rerio (Taxon ID: 7955)
Host Organism:
Method Details:
Experimental Method:
Resolution:
2.30 Å
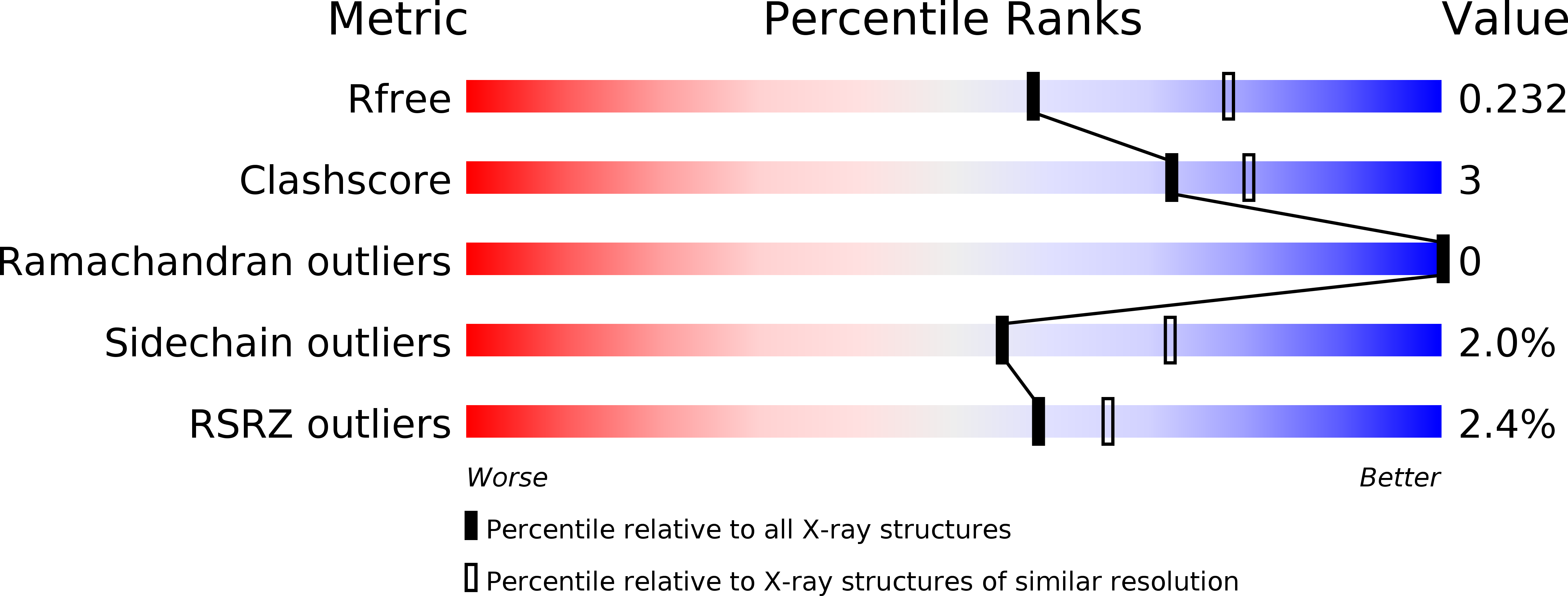
R-Value Free:
0.23
R-Value Work:
0.17
R-Value Observed:
0.17
Space Group:
P 1 21 1