Deposition Date
2019-01-22
Release Date
2019-07-03
Last Version Date
2024-10-16
Entry Detail
PDB ID:
6QJ3
Keywords:
Title:
Crystal structure of the C. thermophilum condensin Ycs4-Brn1 subcomplex
Biological Source:
Source Organism(s):
Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (Taxon ID: 759272)
Chaetomium thermophilum (Taxon ID: 209285)
Chaetomium thermophilum (Taxon ID: 209285)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.30 Å
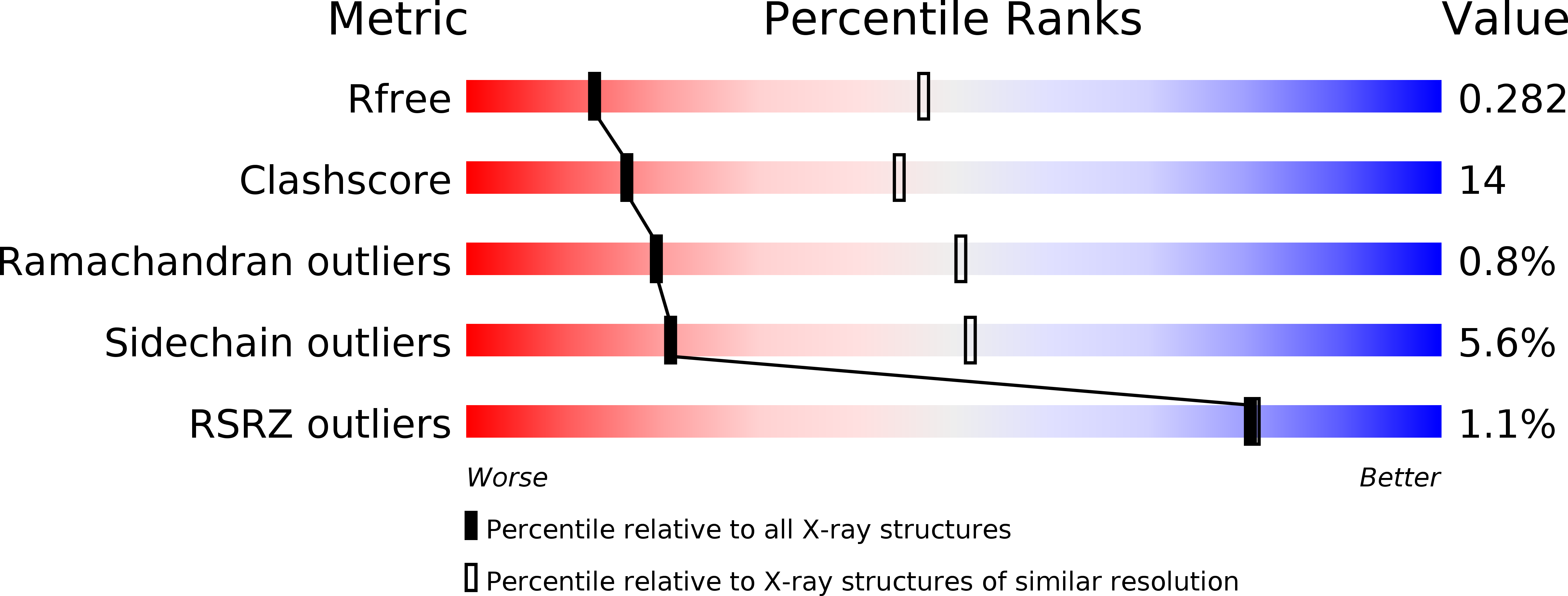
R-Value Free:
0.28
R-Value Work:
0.23
R-Value Observed:
0.23
Space Group:
P 1 21 1