Deposition Date
2019-04-09
Release Date
2019-09-18
Last Version Date
2025-04-02
Entry Detail
PDB ID:
6OIA
Keywords:
Title:
(1S,3S)-3-amino-4-(perfluoropropan-2-ylidene)cyclopentane-1-carboxylic acid hydrochloride, a potent inhibitor of ornithine aminotransferase
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.78 Å
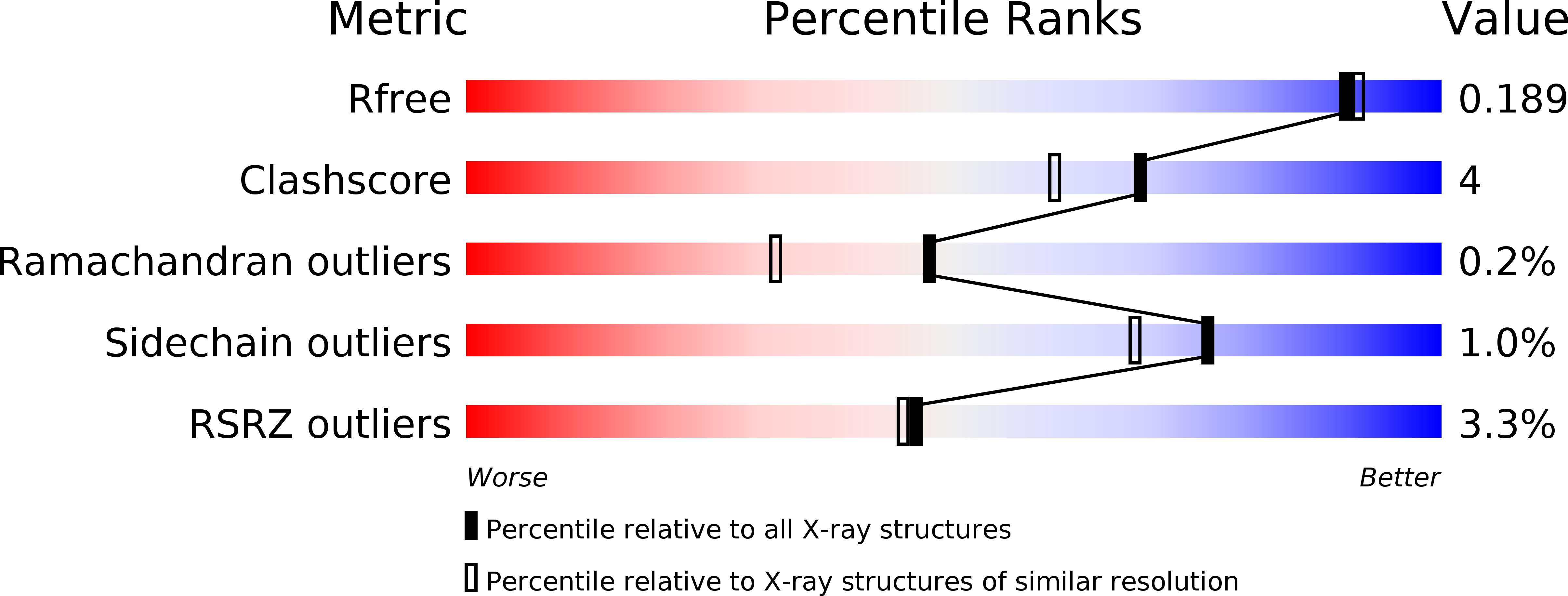
R-Value Free:
0.18
R-Value Work:
0.16
R-Value Observed:
0.16
Space Group:
P 32 2 1


