Deposition Date
2019-01-03
Release Date
2020-01-15
Last Version Date
2025-09-17
Entry Detail
PDB ID:
6J31
Keywords:
Title:
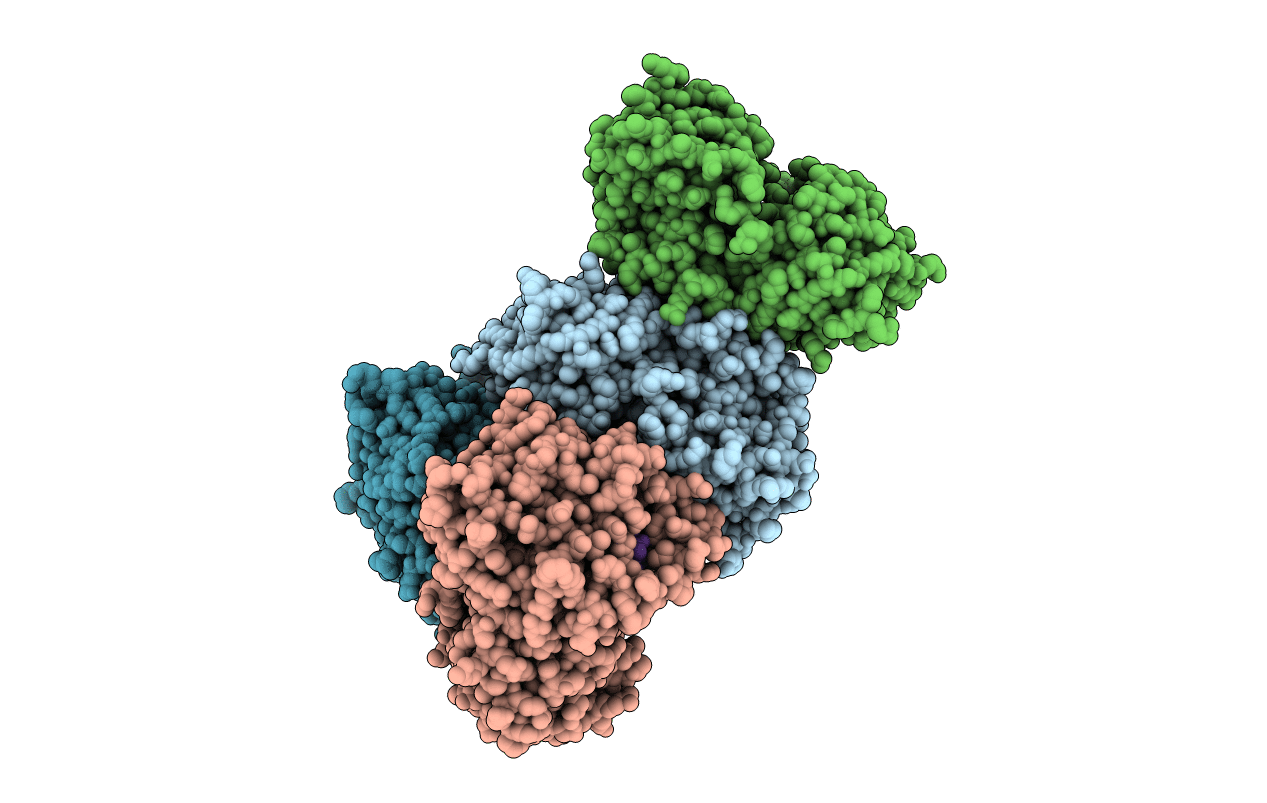
Crystal Structure Analysis of the Glycotransferase of kitacinnamycin
Biological Source:
Source Organism(s):
Kitasatospora (Taxon ID: 2063)
synthetic construct (Taxon ID: 32630)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.24 Å
R-Value Free:
0.24
R-Value Work:
0.22
R-Value Observed:
0.22
Space Group:
I 2 3


