Deposition Date
2017-09-30
Release Date
2017-12-27
Last Version Date
2024-10-23
Entry Detail
PDB ID:
6ELZ
Keywords:
Title:
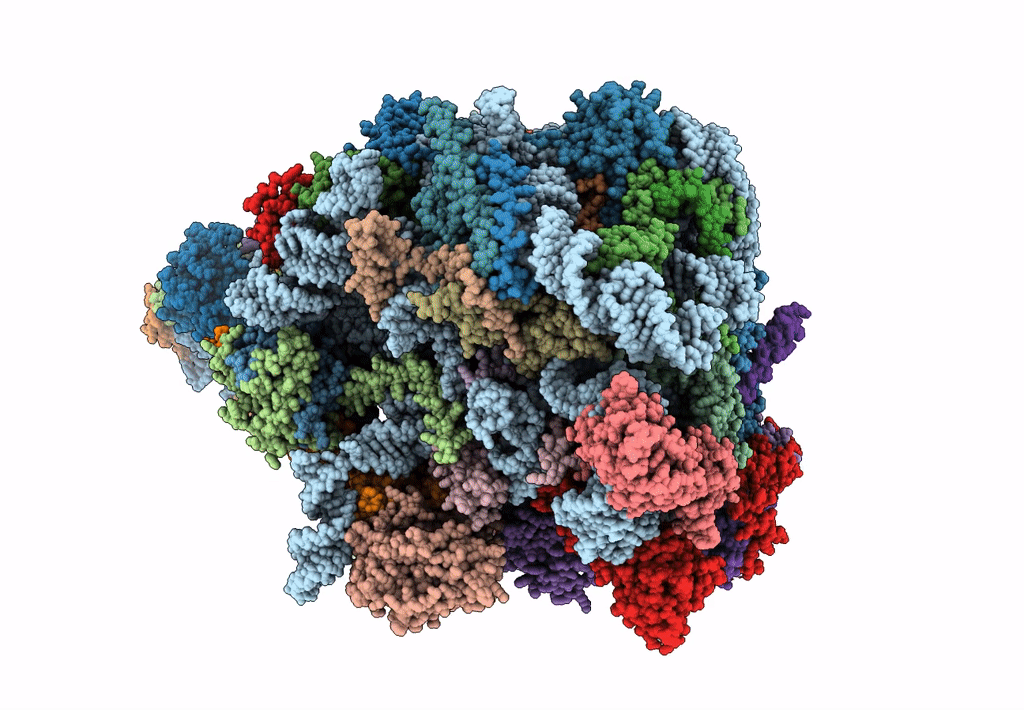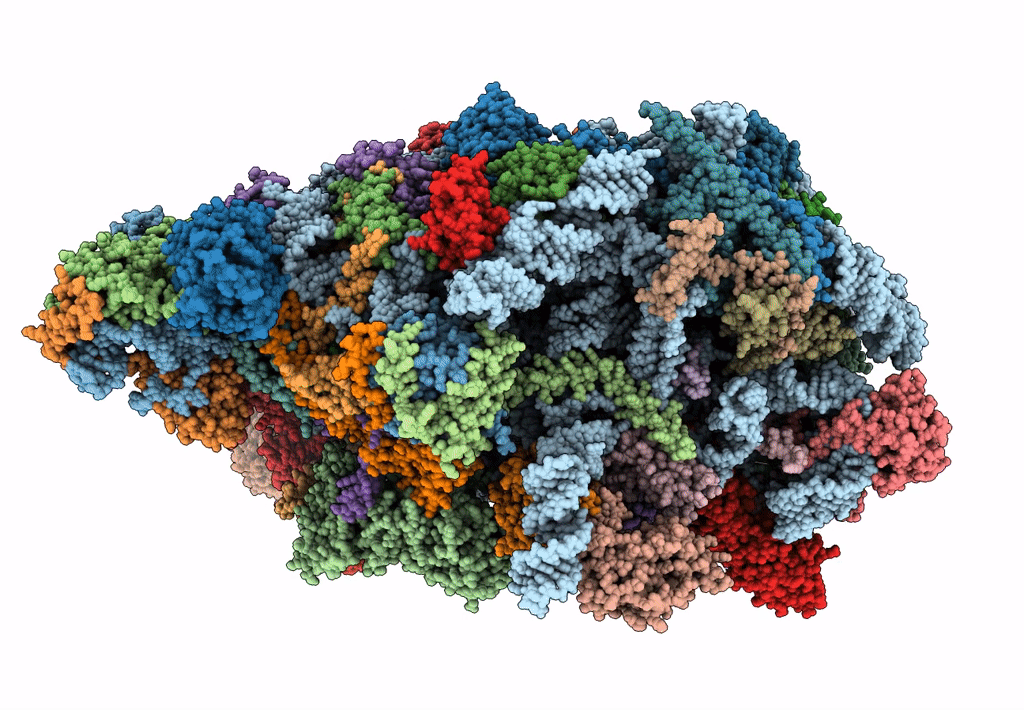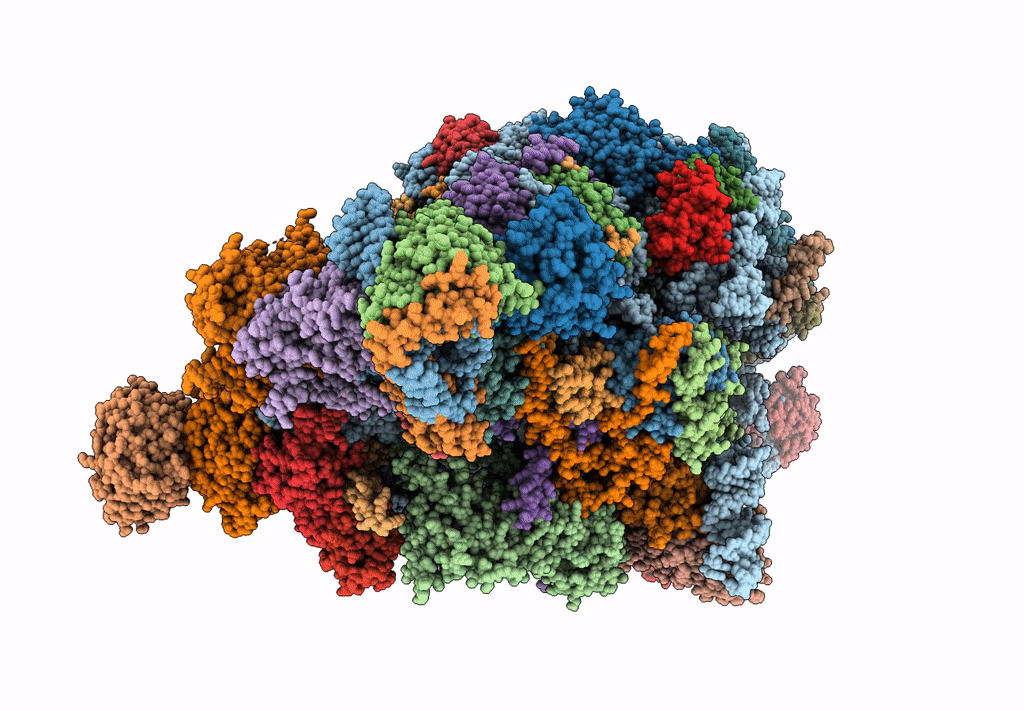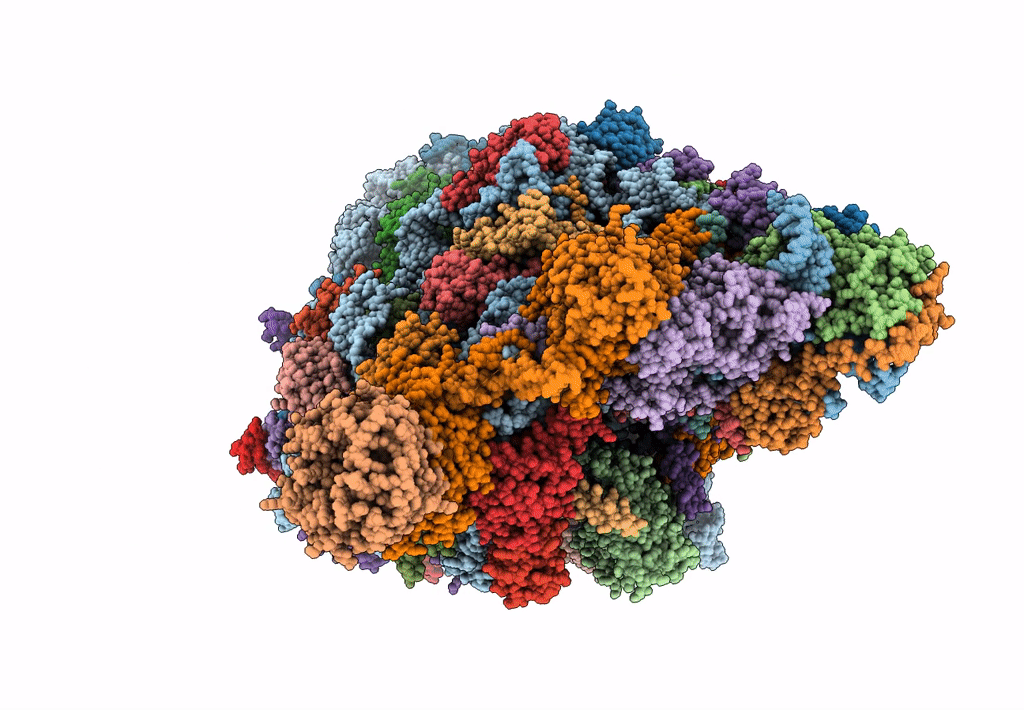
State E (TAP-Flag-Ytm1 E80A) - Visualizing the assembly pathway of nucleolar pre-60S ribosomes
Biological Source:
Source Organism(s):
Saccharomyces cerevisiae S288c (Taxon ID: 559292)
Method Details:
Experimental Method:
Resolution:
3.30 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE


