Deposition Date
2018-07-02
Release Date
2019-03-20
Last Version Date
2024-11-13
Entry Detail
PDB ID:
6DZ2
Keywords:
Title:
Crystal structure of human 5'-deoxy-5'-methylthioadenosine phosphorylase in complex with (3R,4S)-1-((4-amino-5H-pyrrolo[3,2-d]pyrimidin-7-yl)methyl)-4-(((3-(1-benzyl-1H-1,2,3-triazol-4-yl)propyl)thio)methyl)pyrrolidin-3-ol
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.99 Å
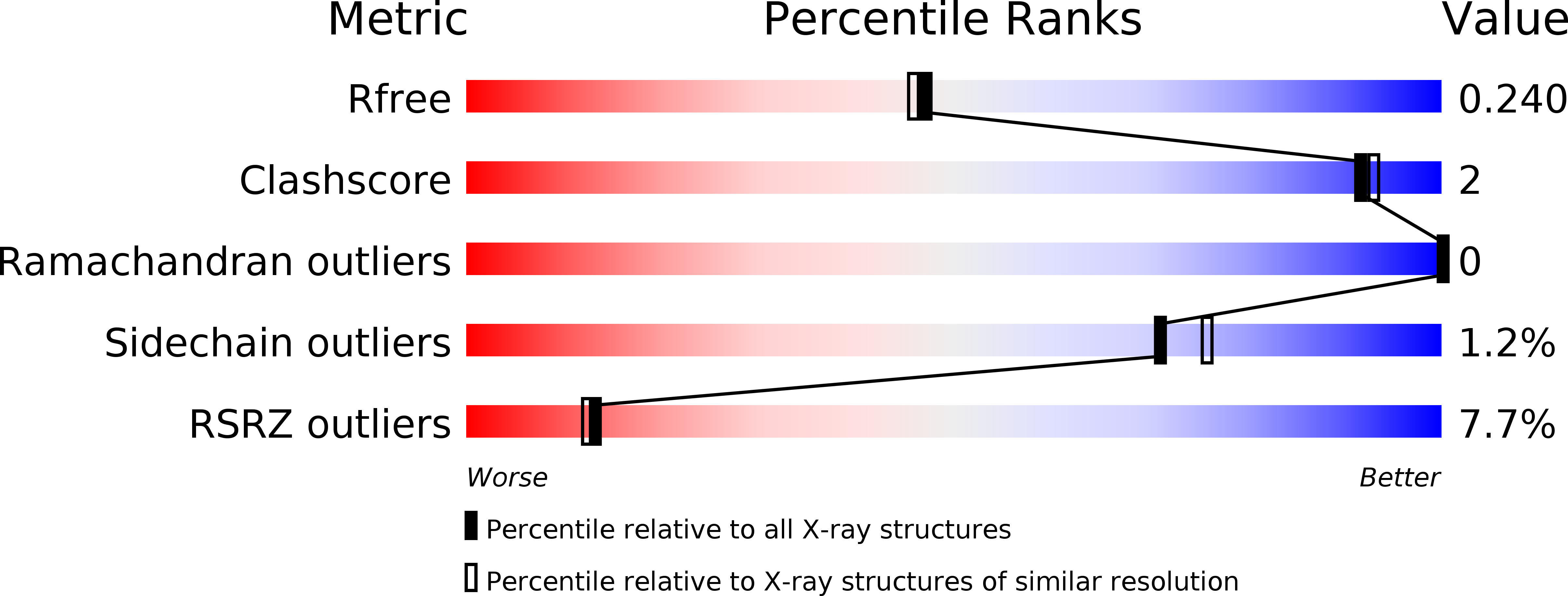
R-Value Free:
0.23
R-Value Work:
0.20
R-Value Observed:
0.20
Space Group:
C 2 2 21