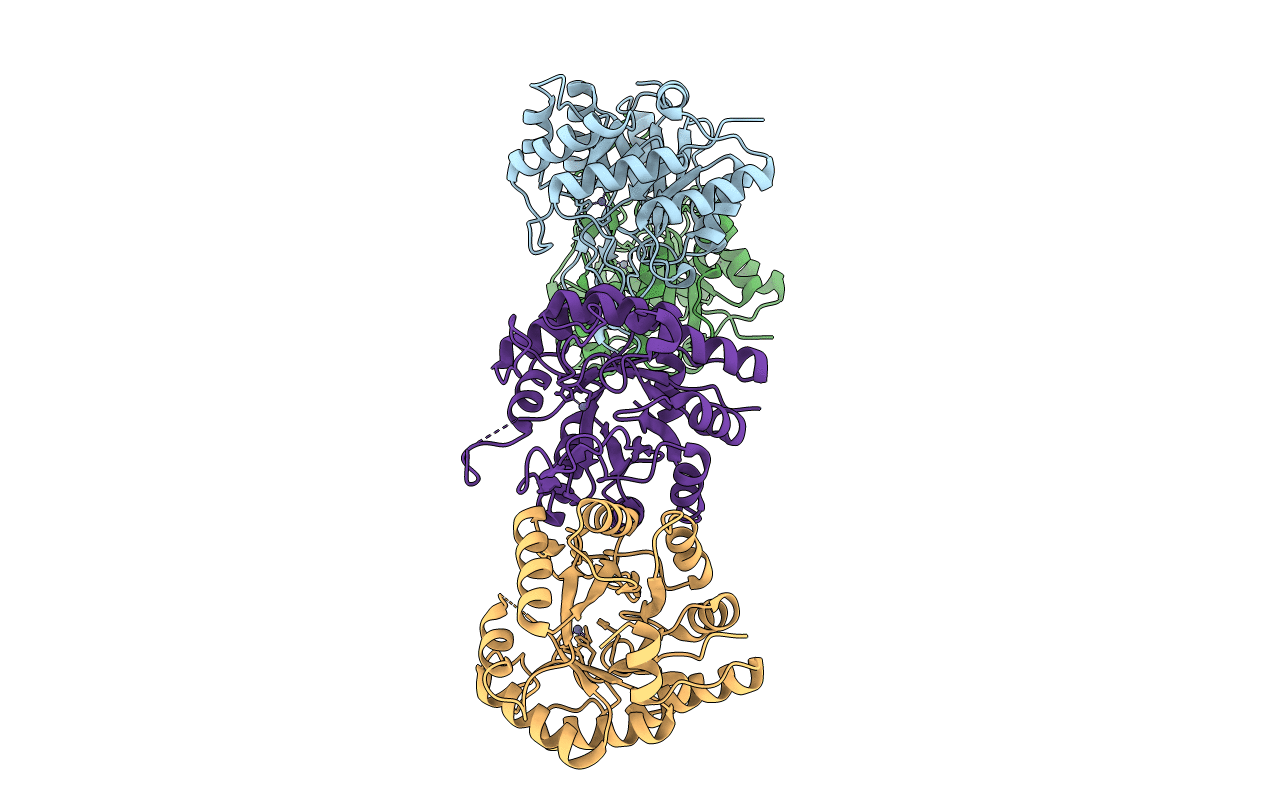
Deposition Date
2018-04-06
Release Date
2018-08-08
Last Version Date
2023-11-22
Entry Detail
PDB ID:
5ZMU
Keywords:
Title:
Crystal structure of a cis-epoxysuccinate hydrolase producing D(-)-tartaric acids
Biological Source:
Source Organism(s):
Bordetella sp. BK-52 (Taxon ID: 454599)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.50 Å
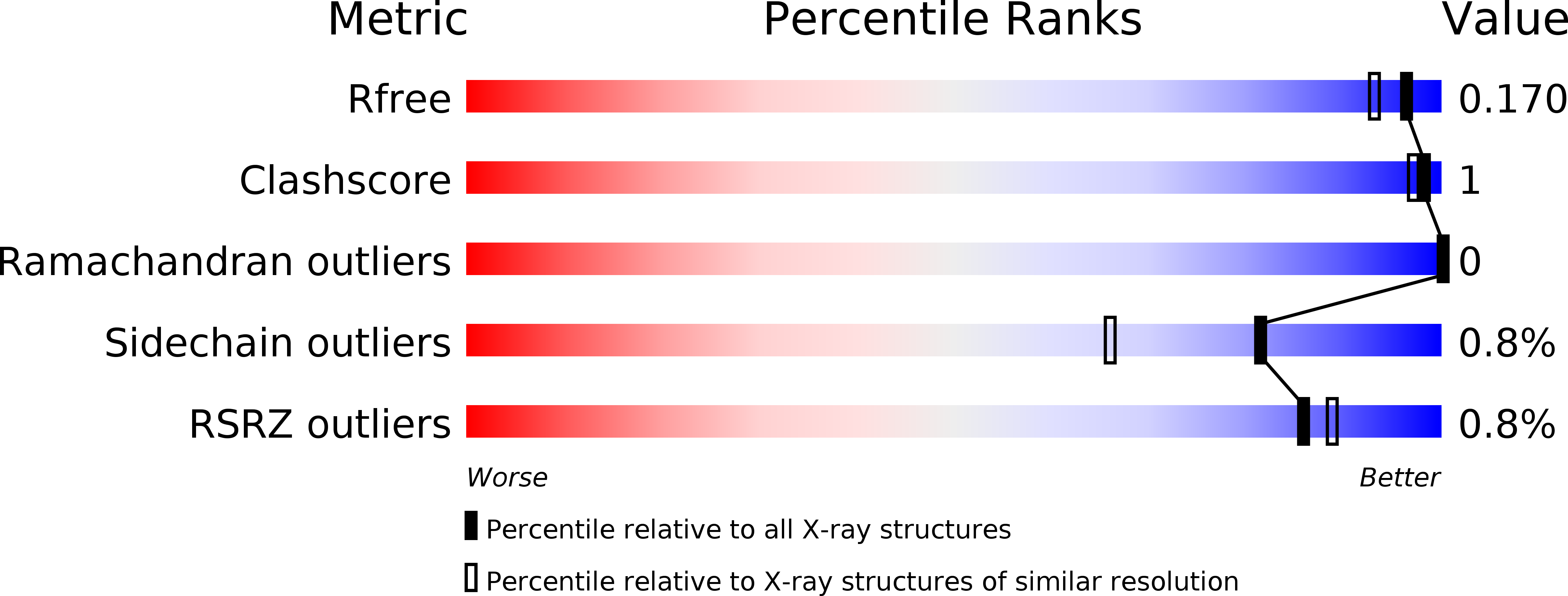
R-Value Free:
0.17
R-Value Work:
0.14
R-Value Observed:
0.14
Space Group:
P 1 2 1


