Deposition Date
2017-02-09
Release Date
2017-04-19
Last Version Date
2024-03-27
Entry Detail
PDB ID:
5X41
Keywords:
Title:
3.5A resolution structure of a cobalt energy-coupling factor transporter using LCP method-CbiMQO
Biological Source:
Source Organism(s):
Rhodobacter capsulatus (Taxon ID: 1061)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.47 Å
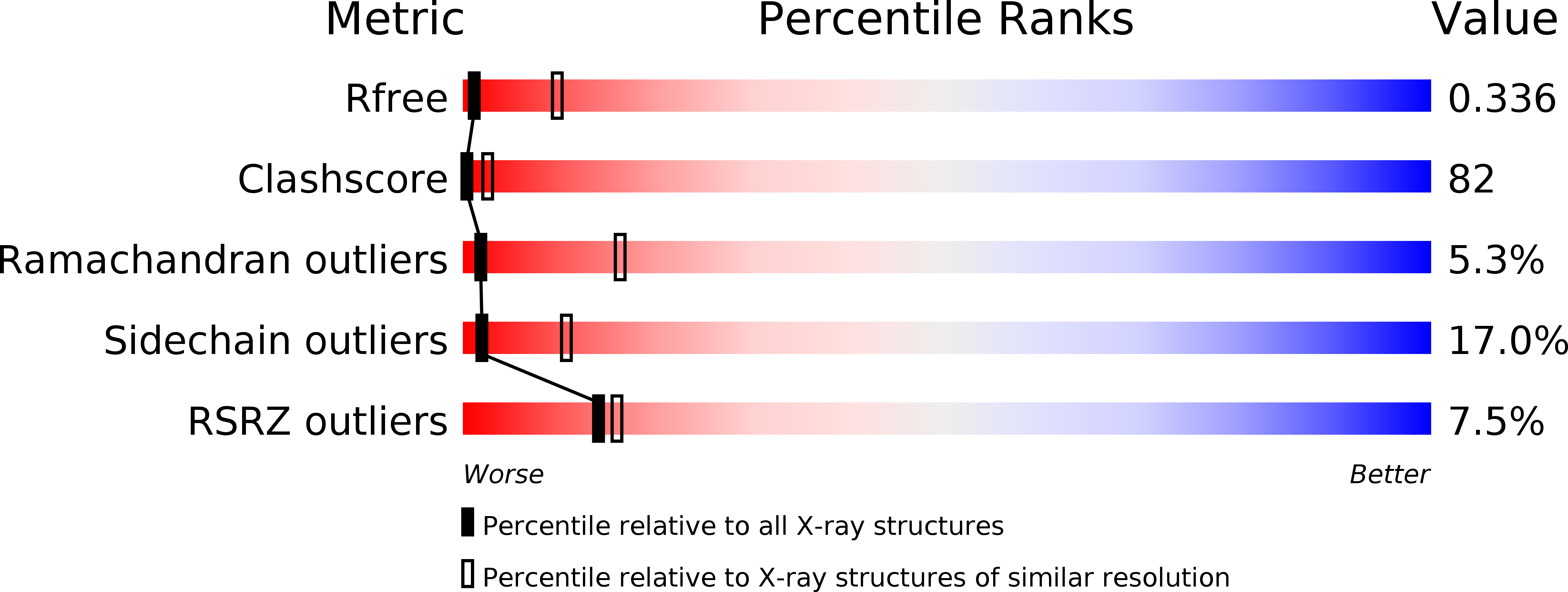
R-Value Free:
0.33
R-Value Work:
0.29
R-Value Observed:
0.30
Space Group:
P 21 21 21