Deposition Date
2017-06-02
Release Date
2018-06-06
Last Version Date
2024-03-13
Entry Detail
PDB ID:
5W1A
Keywords:
Title:
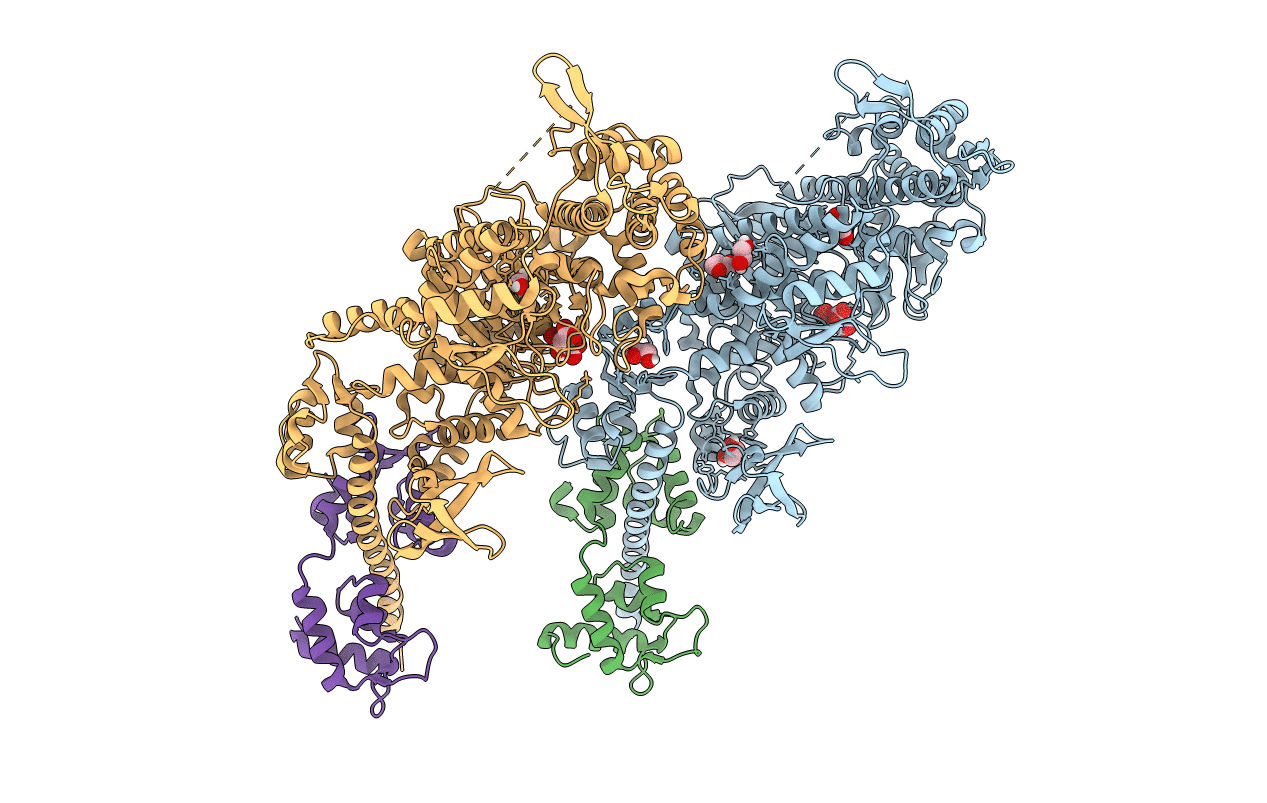
The first X-ray crystal structure of an insect muscle myosin. Drosophila melanogaster, skeletal muscle myosin II, an embryonic isoform, subfragment-1
Biological Source:
Source Organism:
Drosophila melanogaster (Taxon ID: 7227)
Host Organism:
Method Details:
Experimental Method:
Resolution:
2.23 Å
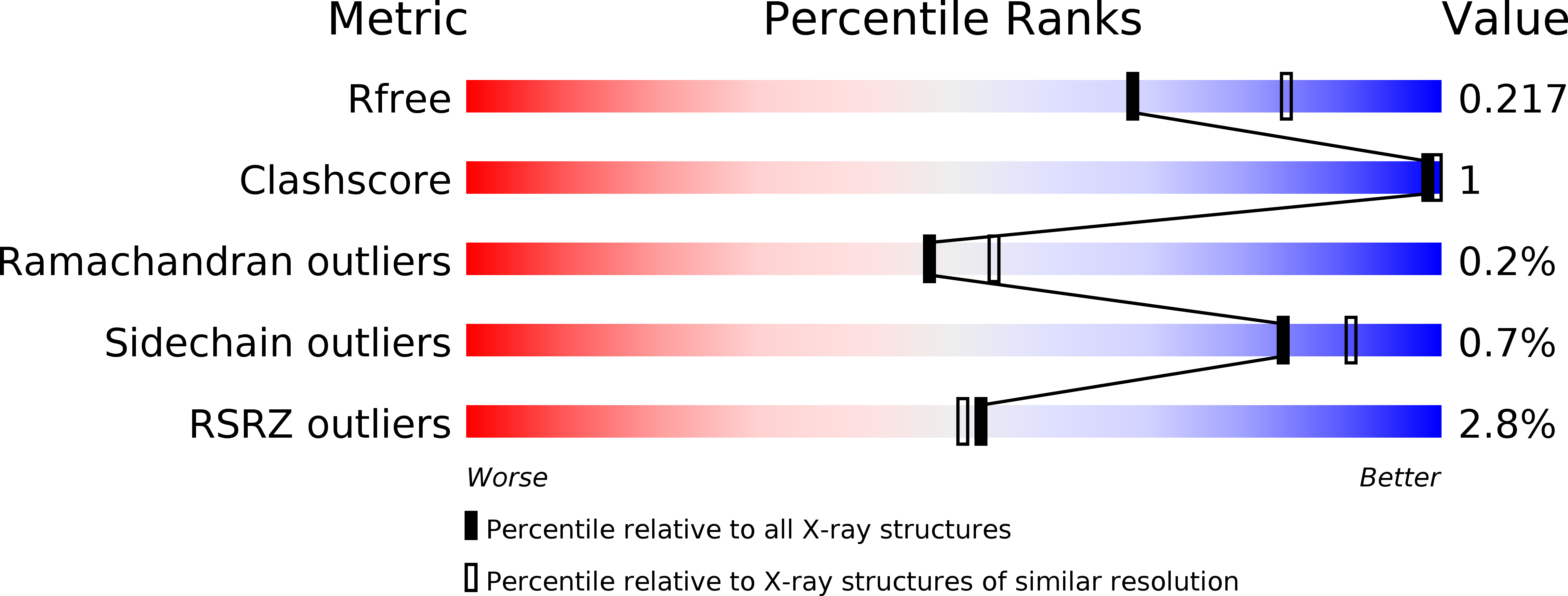
R-Value Free:
0.21
R-Value Work:
0.17
R-Value Observed:
0.18
Space Group:
P 21 21 21


