Deposition Date
2017-04-26
Release Date
2018-05-02
Last Version Date
2024-11-20
Entry Detail
PDB ID:
5VM0
Keywords:
Title:
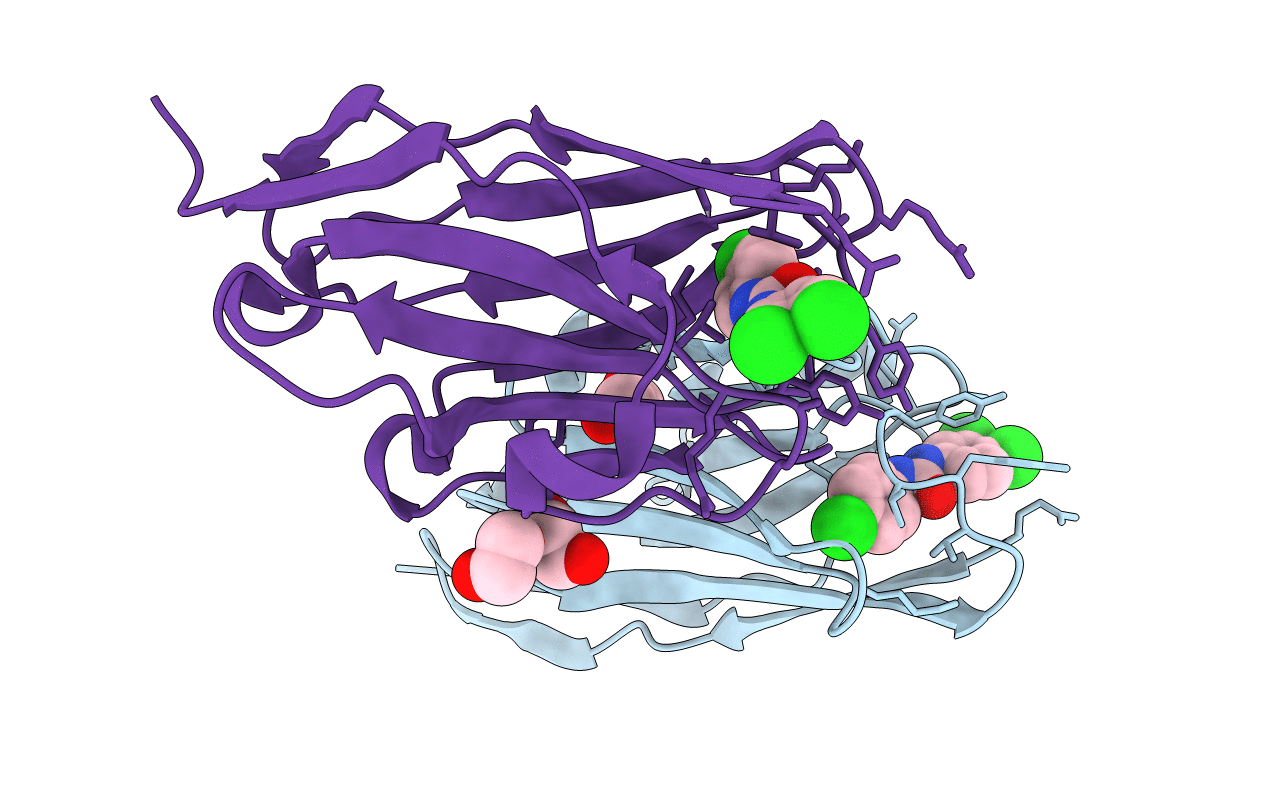
The hapten triclocarban bound to the single domain camelid nanobody VHH T9
Biological Source:
Source Organism:
Lama glama (Taxon ID: 9844)
Host Organism:
Method Details:
Experimental Method:
Resolution:
1.79 Å
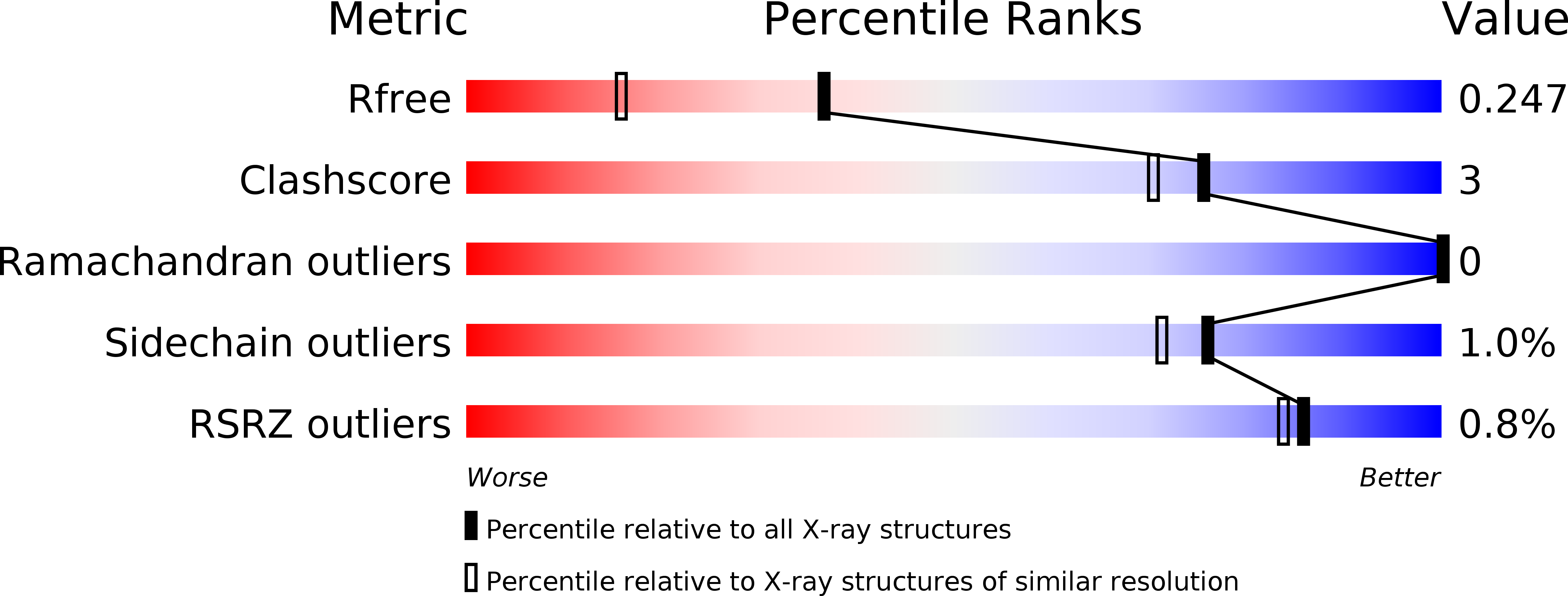
R-Value Free:
0.24
R-Value Work:
0.20
R-Value Observed:
0.20
Space Group:
C 1 2 1