Deposition Date
2016-09-27
Release Date
2017-04-12
Last Version Date
2024-10-23
Entry Detail
PDB ID:
5TG9
Keywords:
Title:
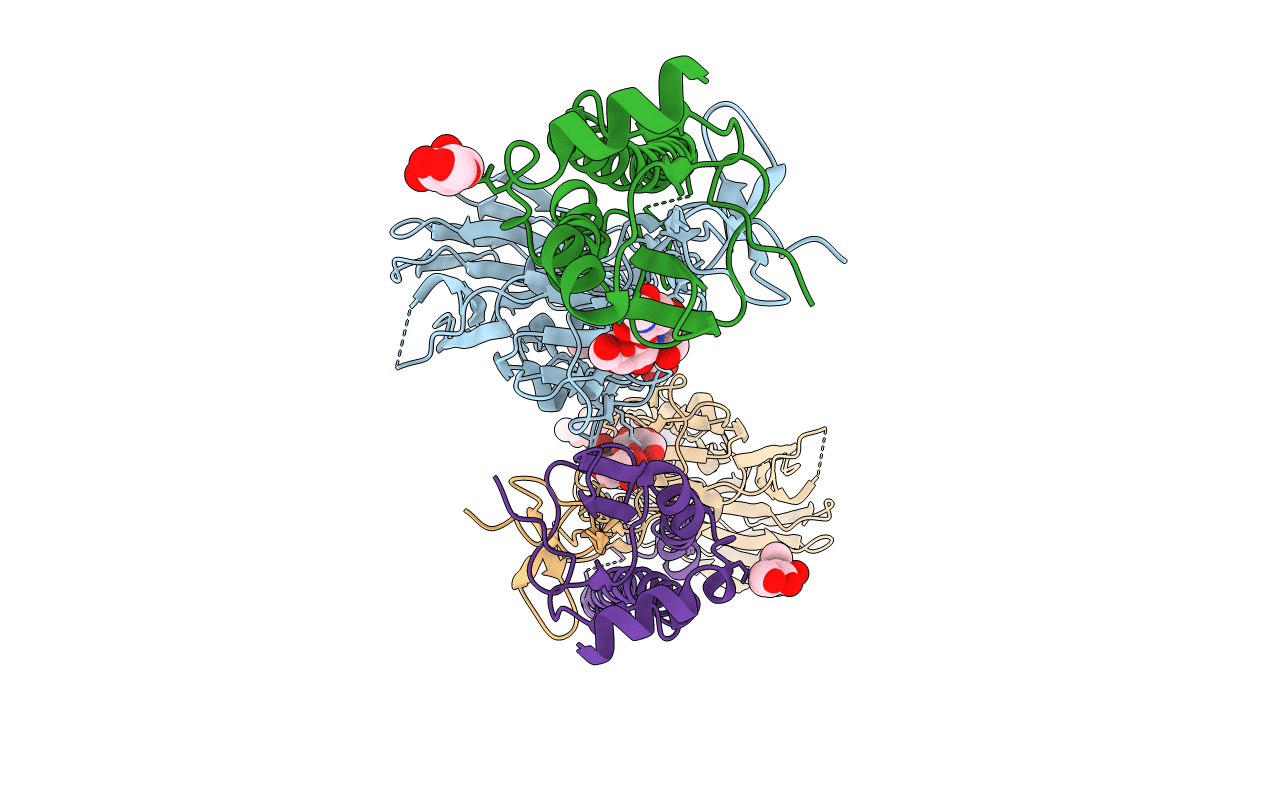
Crystal structure of H15 hemagglutinin from A/shearwater/WA/2576/1979 H15N9 influenza virus in complex with 3'-SLN
Biological Source:
Source Organism(s):
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.75 Å
R-Value Free:
0.27
R-Value Work:
0.22
R-Value Observed:
0.22
Space Group:
P 3


