Deposition Date
2017-08-11
Release Date
2018-04-25
Last Version Date
2024-05-15
Entry Detail
PDB ID:
5OQJ
Keywords:
Title:
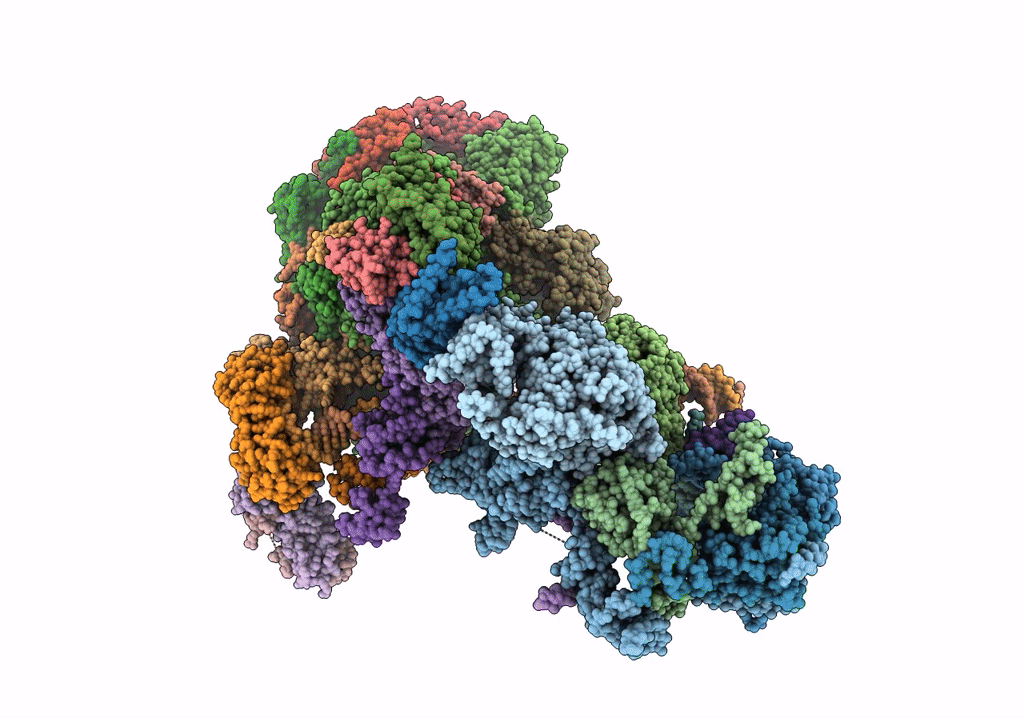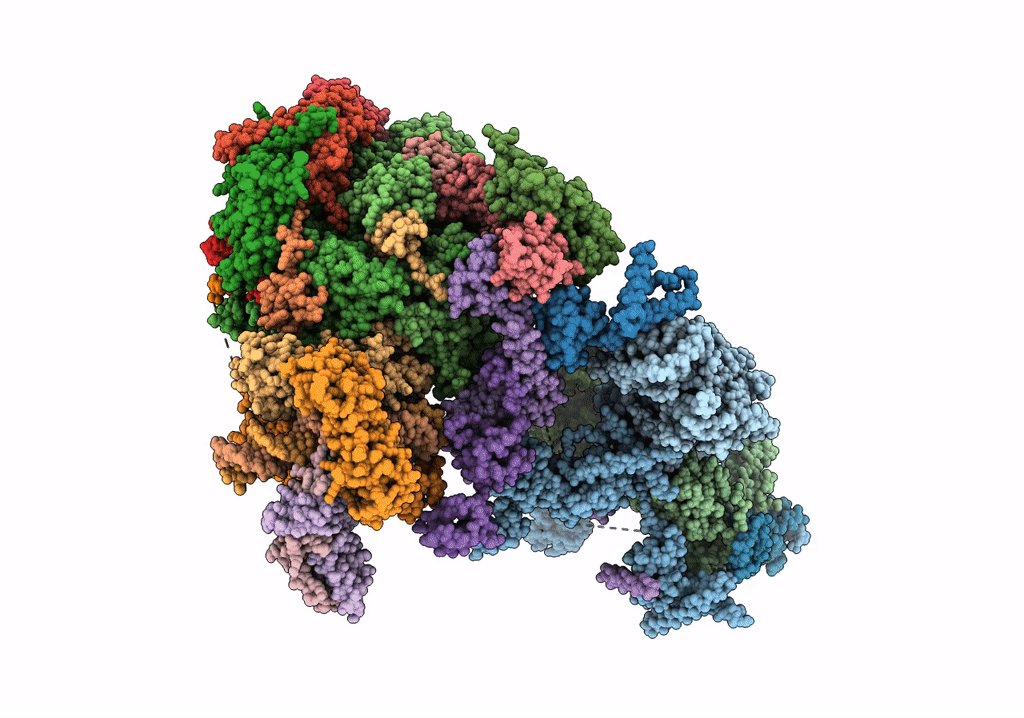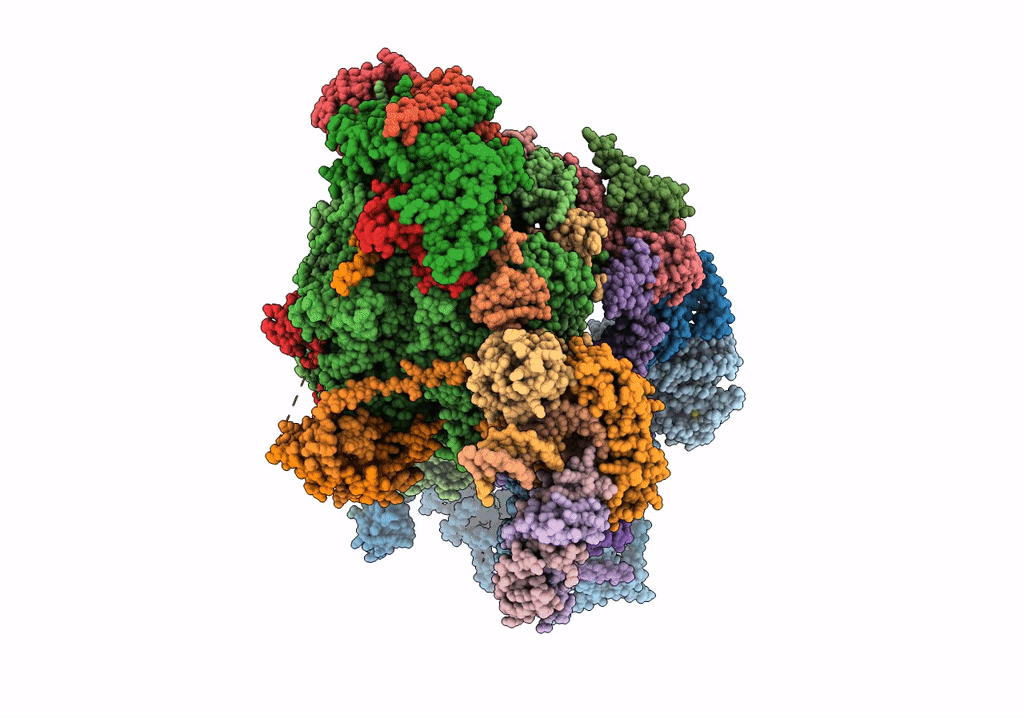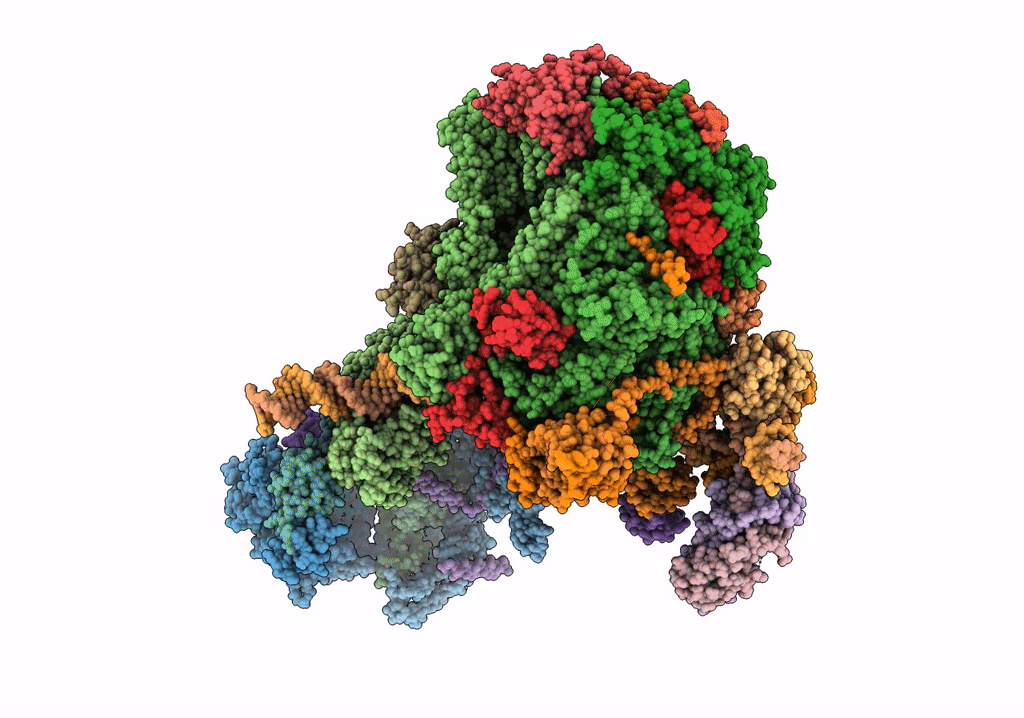
STRUCTURE OF YEAST TRANSCRIPTION PRE-INITIATION COMPLEX WITH TFIIH
Biological Source:
Source Organism(s):
Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (Taxon ID: 559292)
synthetic construct (Taxon ID: 32630)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
4.70 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE