Deposition Date
2015-05-17
Release Date
2016-04-06
Last Version Date
2024-10-16
Entry Detail
PDB ID:
4ZUT
Keywords:
Title:
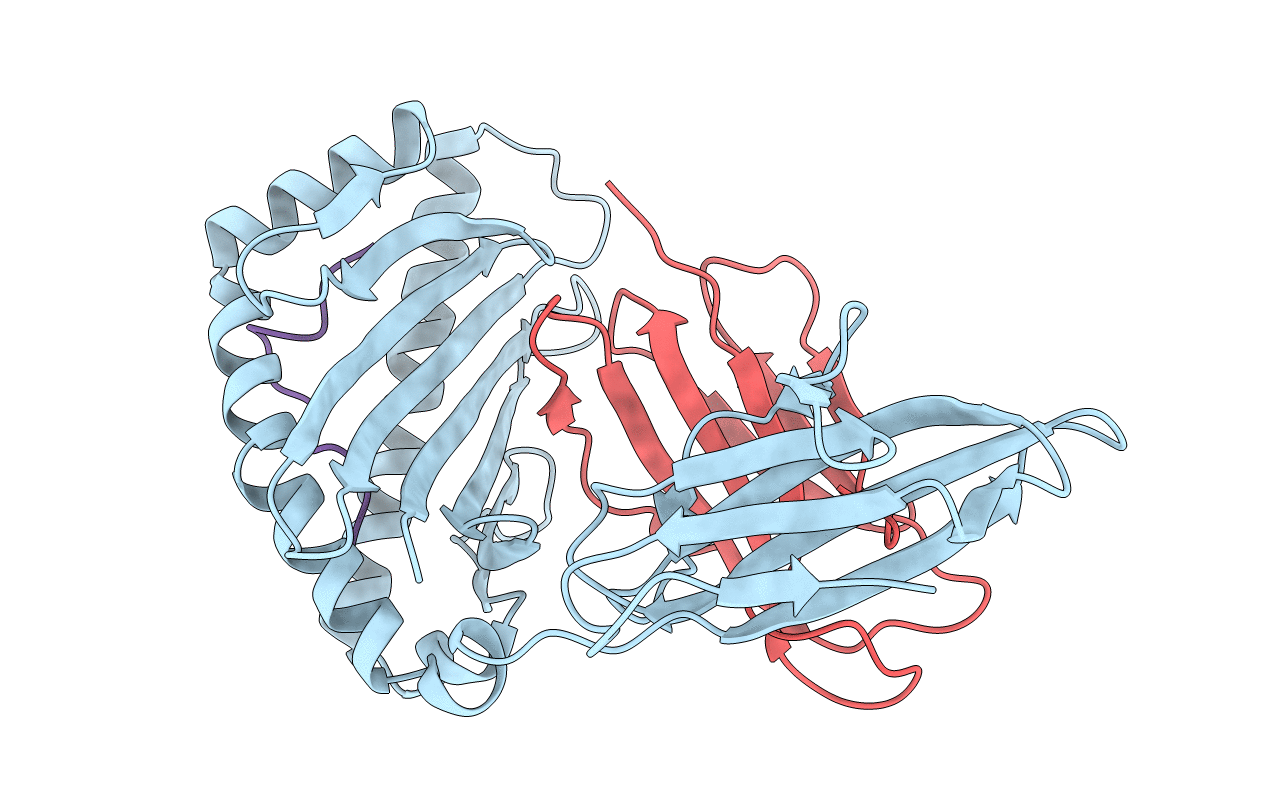
Crystal structure of Equine MHC I(Eqca-N*00602) in complexed with equine infectious anaemia virus (EIAV) derived peptide Gag-GW12
Biological Source:
Source Organism(s):
Equus caballus (Taxon ID: 9796)
Mus musculus (Taxon ID: 10090)
Equine infectious anemia virus (Taxon ID: 11665)
Mus musculus (Taxon ID: 10090)
Equine infectious anemia virus (Taxon ID: 11665)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.60 Å
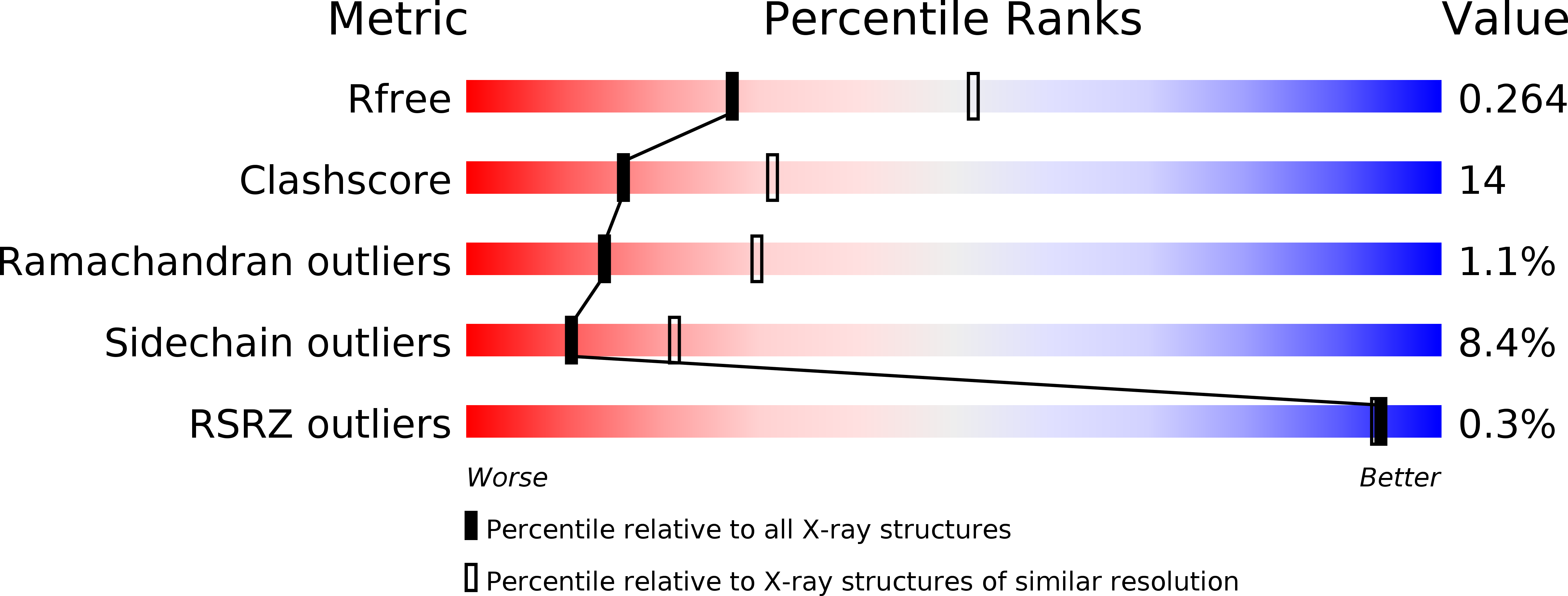
R-Value Free:
0.26
R-Value Work:
0.22
R-Value Observed:
0.22
Space Group:
P 41 21 2