Deposition Date
2015-03-25
Release Date
2015-10-21
Last Version Date
2023-09-27
Entry Detail
PDB ID:
4YZV
Keywords:
Title:
Precleavage 70S structure of the P. vulgaris HigB deltaH92 toxin bound to the ACA codon
Biological Source:
Source Organism:
Proteus vulgaris (Taxon ID: 585)
Escherichia coli (Taxon ID: 562)
Streptomyces alboniger (Taxon ID: 132473)
Thermus thermophilus HB8 (Taxon ID: 300852)
Escherichia coli (Taxon ID: 562)
Streptomyces alboniger (Taxon ID: 132473)
Thermus thermophilus HB8 (Taxon ID: 300852)
Host Organism:
Method Details:
Experimental Method:
Resolution:
3.10 Å
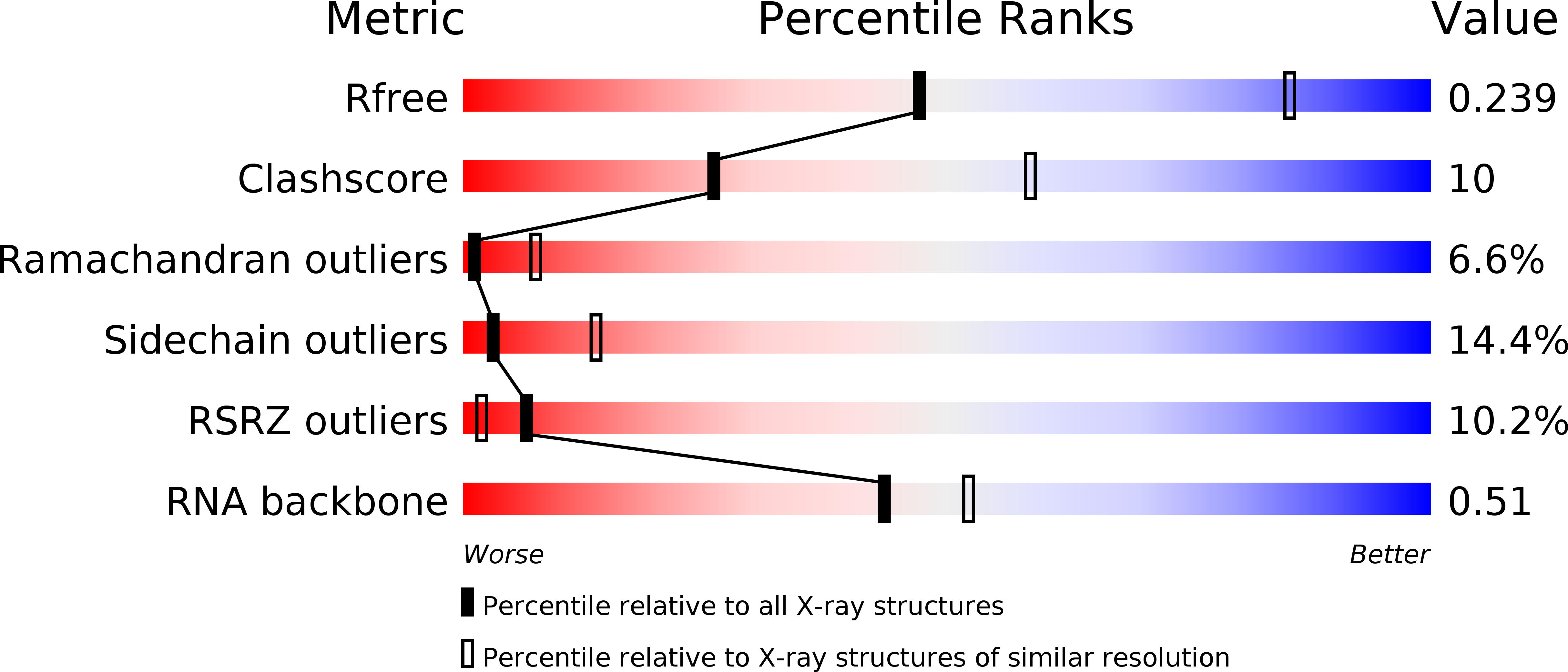
R-Value Free:
0.23
R-Value Work:
0.20
R-Value Observed:
0.20
Space Group:
P 21 21 21