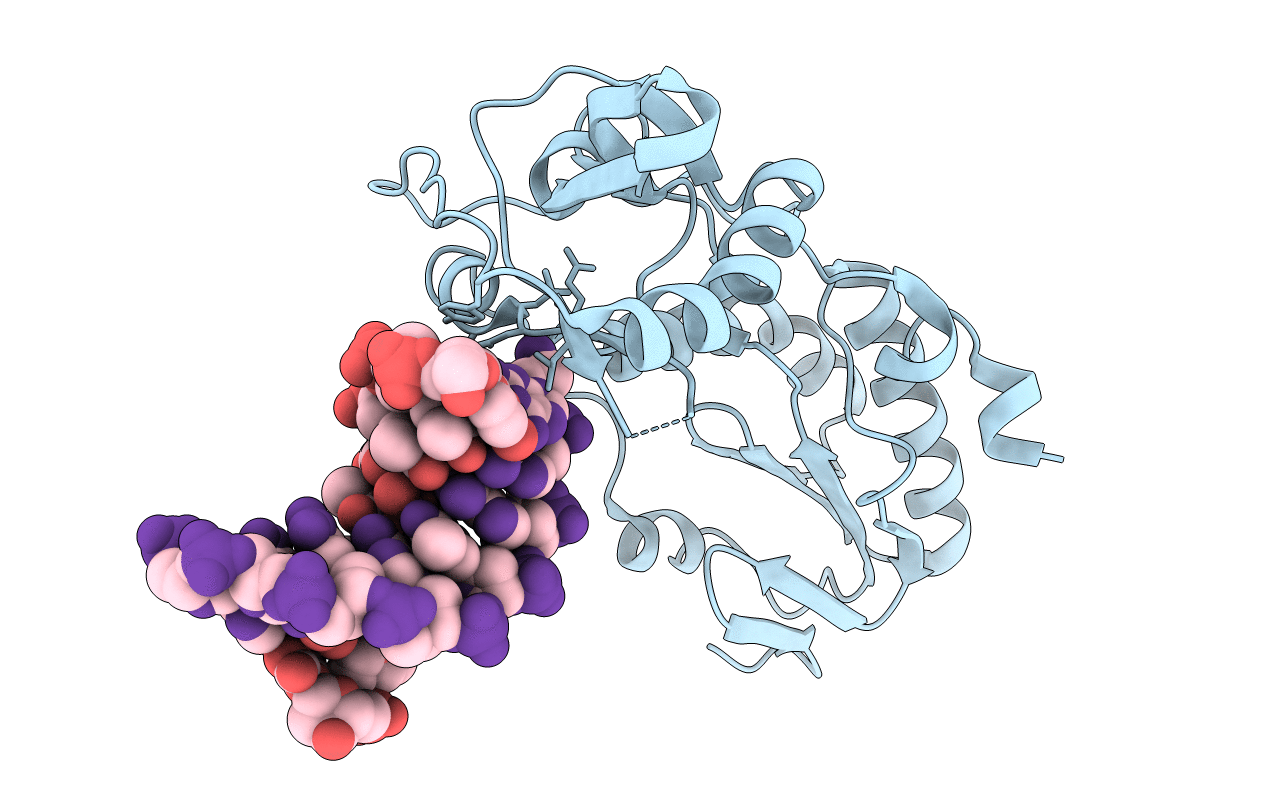
Deposition Date
2015-01-16
Release Date
2015-05-27
Last Version Date
2024-02-28
Entry Detail
PDB ID:
4XO0
Keywords:
Title:
Crystal structure of 5'-CTTATPPTAZZATAAG in a host-guest complex
Biological Source:
Source Organism(s):
Moloney murine leukemia virus (isolate Shinnick) (Taxon ID: 928306)
synthetic construct (Taxon ID: 32630)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.70 Å
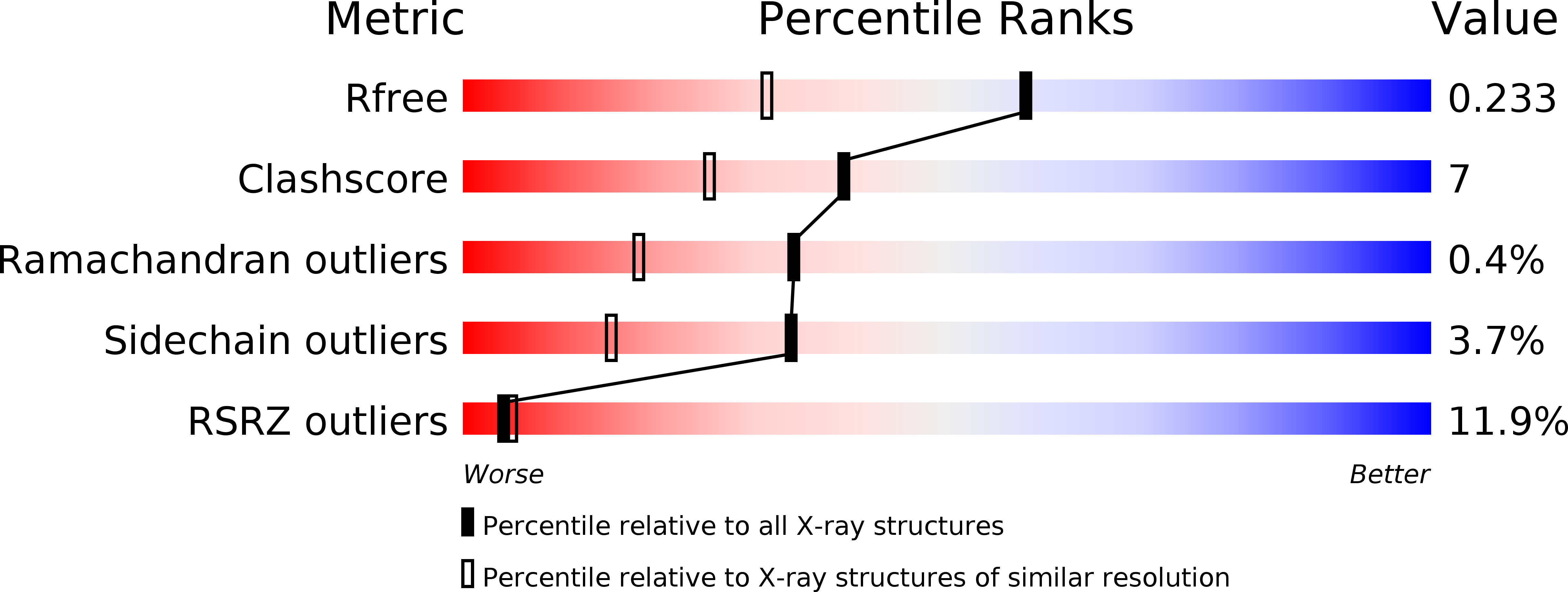
R-Value Free:
0.24
R-Value Work:
0.21
R-Value Observed:
0.22
Space Group:
P 21 21 2