Deposition Date
2014-05-16
Release Date
2014-10-08
Last Version Date
2024-01-10
Entry Detail
PDB ID:
4UMB
Keywords:
Title:
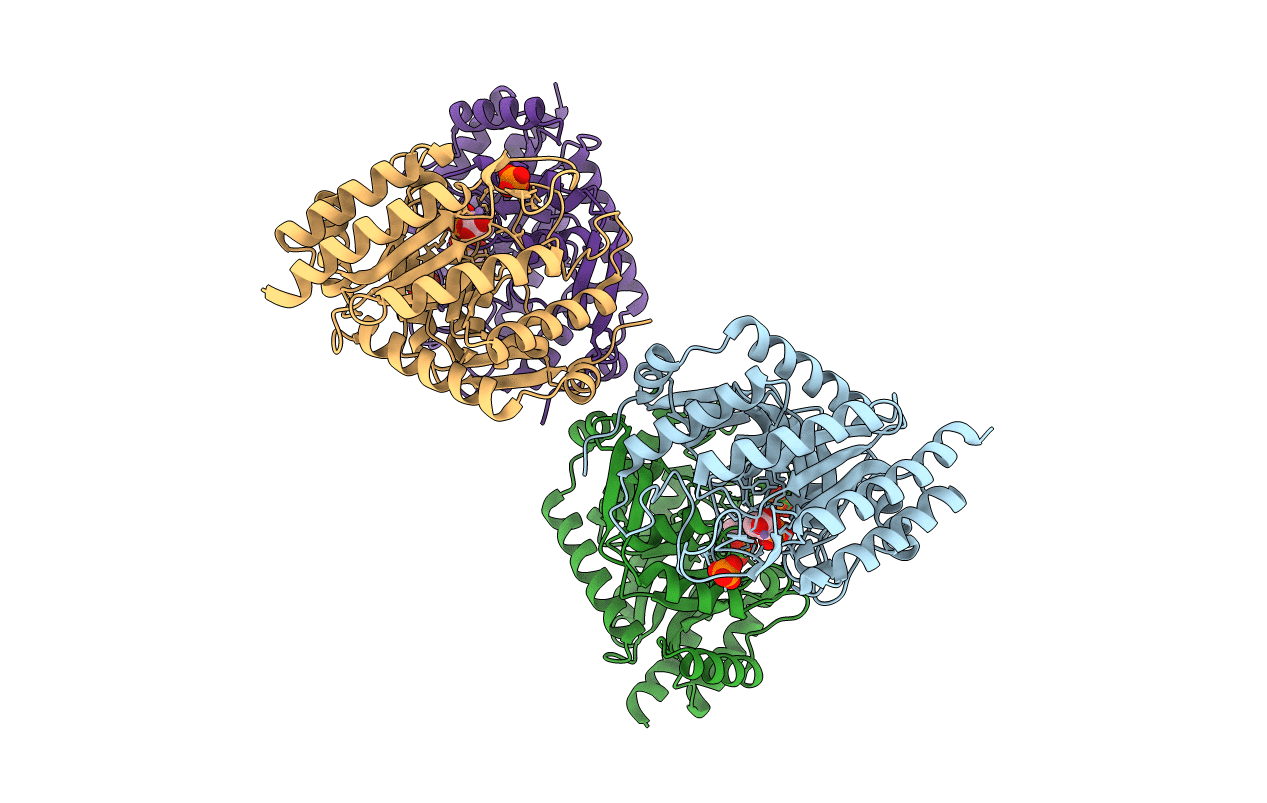
Structural analysis of substrate-mimicking inhibitors in complex with Neisseria meningitidis 3-deoxy-D-arabino-heptulosonate 7-phosphate synthase - the importance of accommodating the active site water
Biological Source:
Source Organism(s):
NEISSERIA MENINGITIDIS (Taxon ID: 122586)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.17 Å
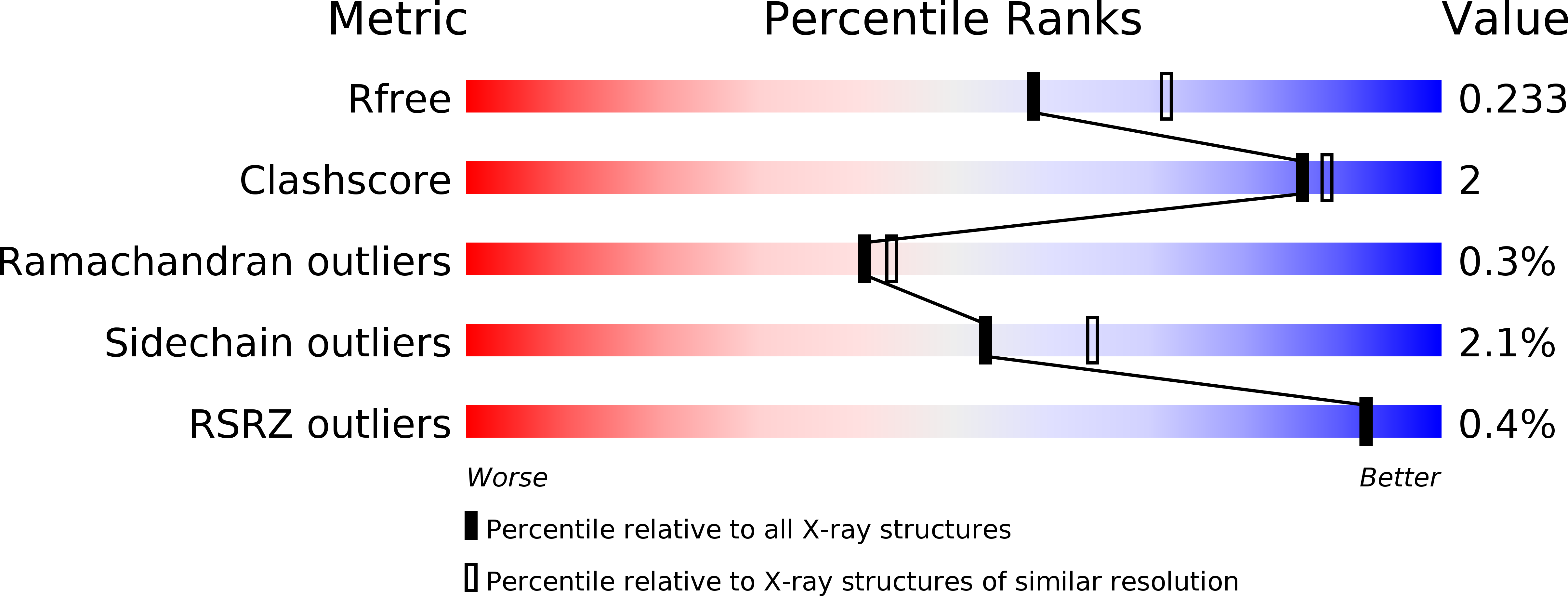
R-Value Free:
0.22
R-Value Work:
0.20
R-Value Observed:
0.20
Space Group:
P 21 21 21