Deposition Date
2014-09-27
Release Date
2015-10-07
Last Version Date
2023-11-08
Entry Detail
PDB ID:
4RFT
Keywords:
Title:
T=1 subviral particle of Grouper nervous necrosis virus capsid protein deletion mutant (delta 1-34 & 218-338)
Biological Source:
Source Organism(s):
Epinephelus coioides nervous necrosis virus (Taxon ID: 204929)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.10 Å
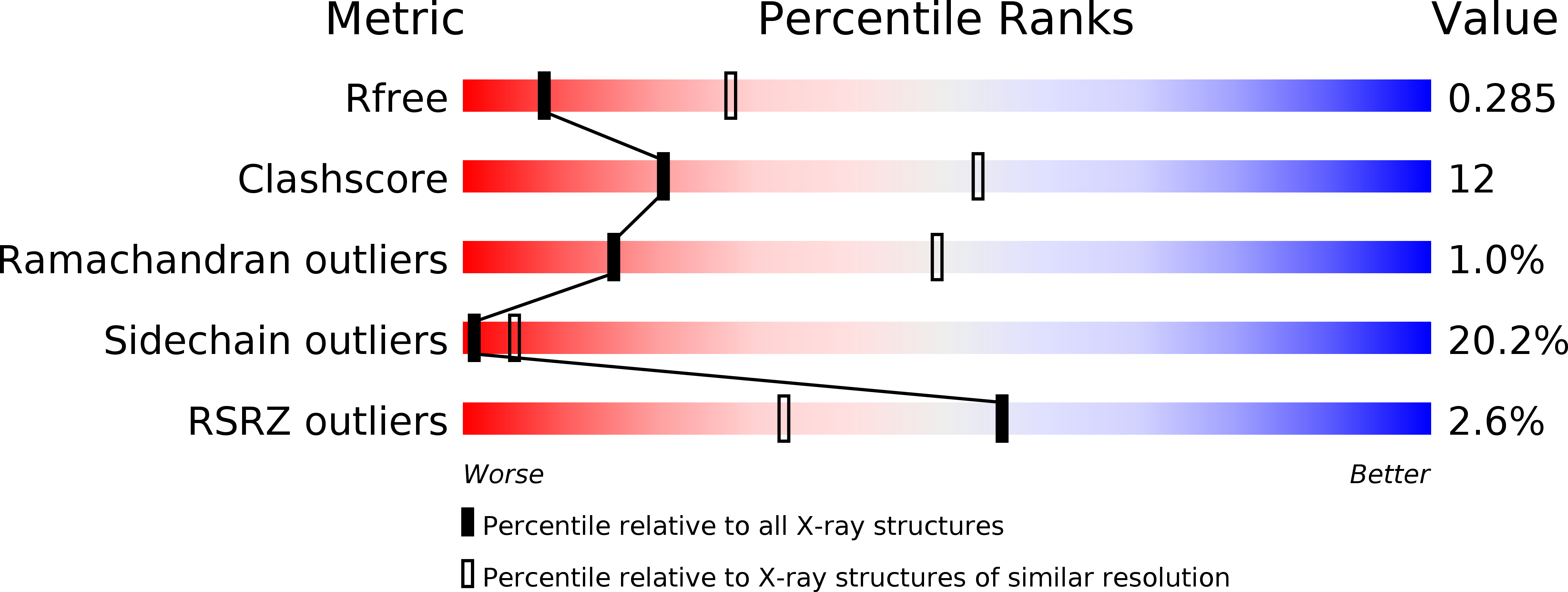
R-Value Free:
0.28
R-Value Work:
0.23
R-Value Observed:
0.24
Space Group:
P 42