Deposition Date
2013-04-04
Release Date
2013-06-26
Last Version Date
2024-10-09
Entry Detail
PDB ID:
4K0K
Keywords:
Title:
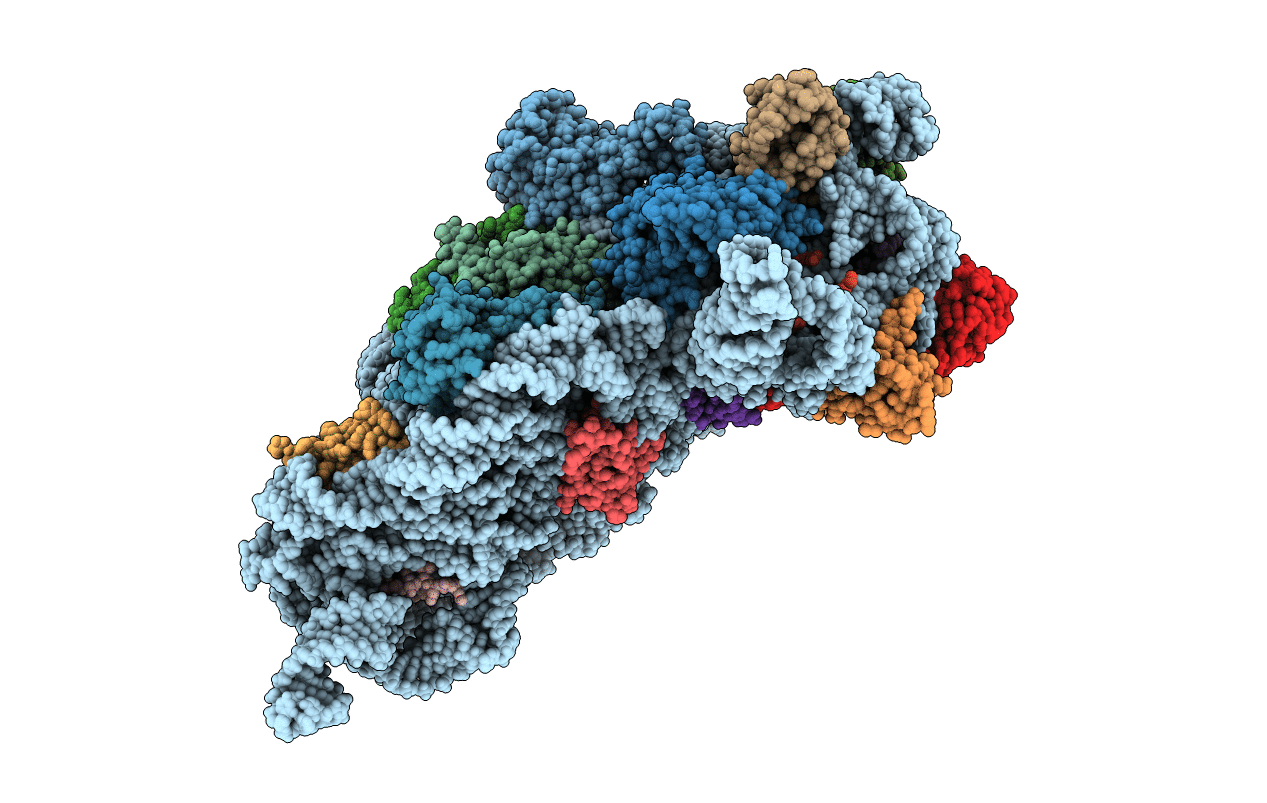
Crystal structure of the Thermus thermophilus 30S ribosomal subunit complexed with a serine-ASL and mRNA containing a stop codon
Biological Source:
Source Organism(s):
Synthetic construct (Taxon ID: 32630)
Thermus thermophilus (Taxon ID: 300852)
Thermus thermophilus (Taxon ID: 300852)
Method Details:
Experimental Method:
Resolution:
3.40 Å
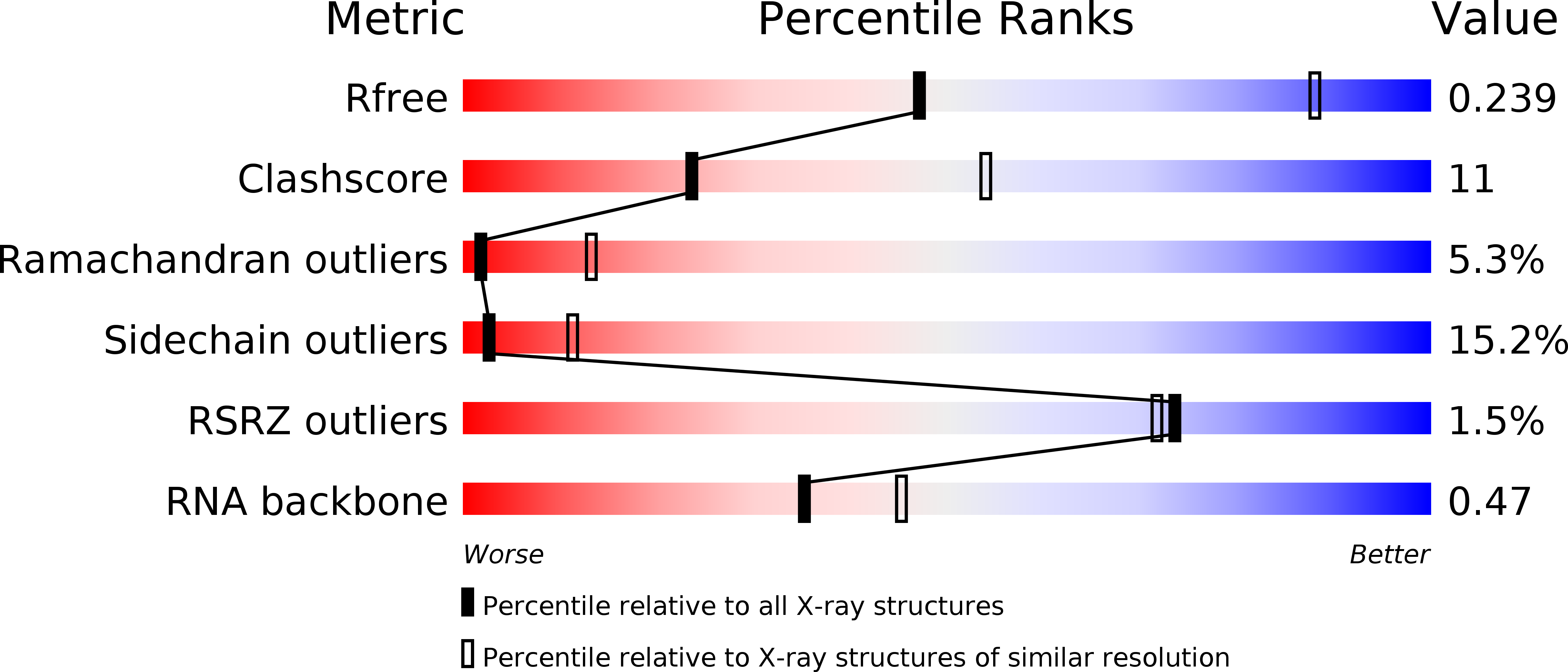
R-Value Free:
0.24
R-Value Work:
0.18
R-Value Observed:
0.19
Space Group:
P 41 21 2