Deposition Date
2014-01-17
Release Date
2014-10-22
Last Version Date
2024-10-16
Entry Detail
PDB ID:
4CMR
Keywords:
Title:
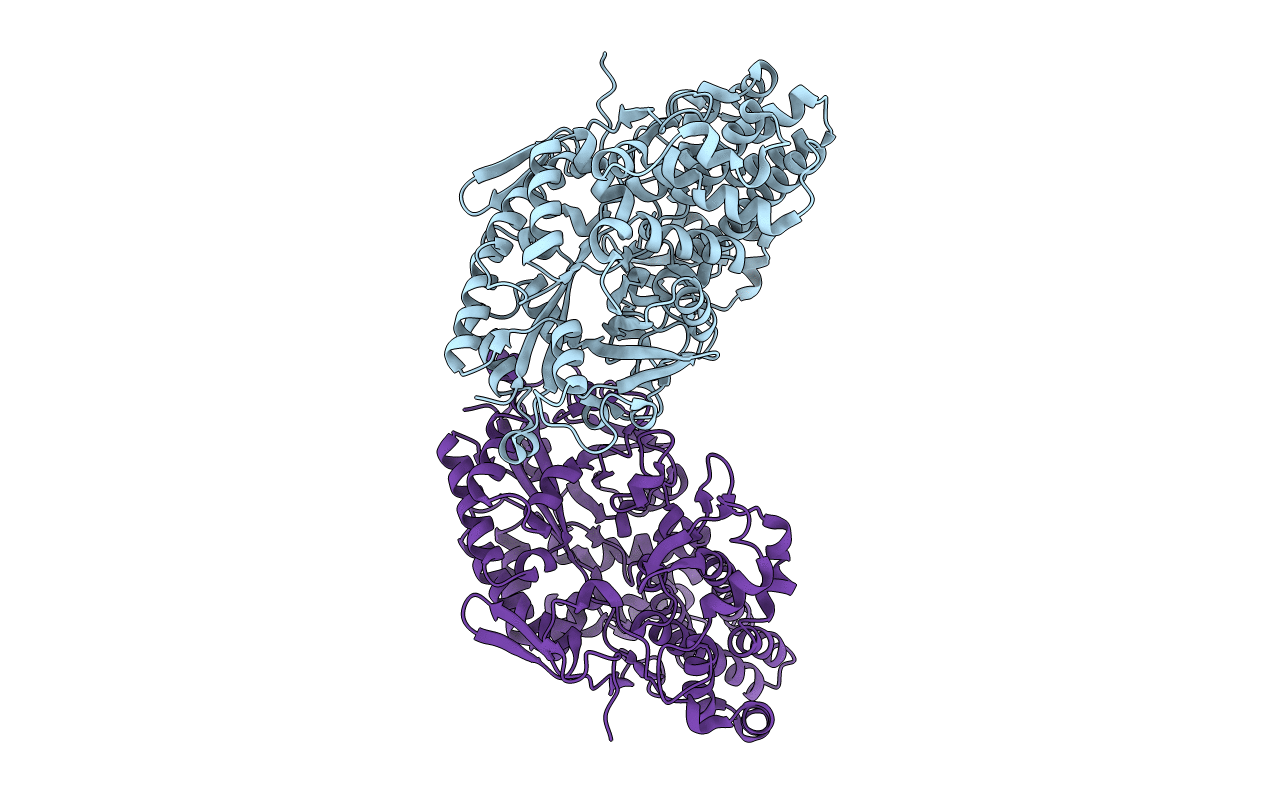
The crystal structure of novel exo-type maltose-forming amylase(Py04_0872) from Pyrococcus sp. ST04
Biological Source:
Source Organism(s):
PYROCOCCUS SP. ST04 (Taxon ID: 1183377)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.80 Å
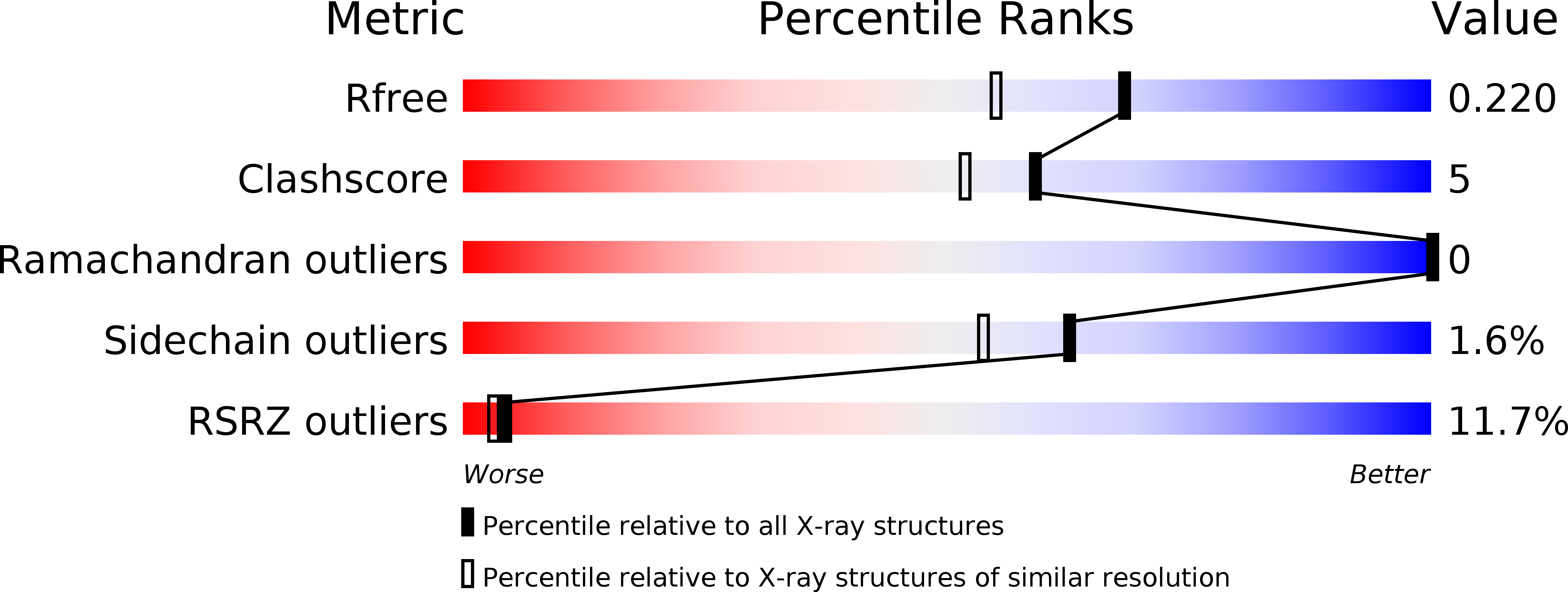
R-Value Free:
0.22
R-Value Work:
0.19
R-Value Observed:
0.19
Space Group:
C 2 2 21