Deposition Date
2013-01-25
Release Date
2013-03-20
Last Version Date
2024-05-08
Entry Detail
Biological Source:
Source Organism(s):
Expression System(s):
Method Details:
Experimental Method:
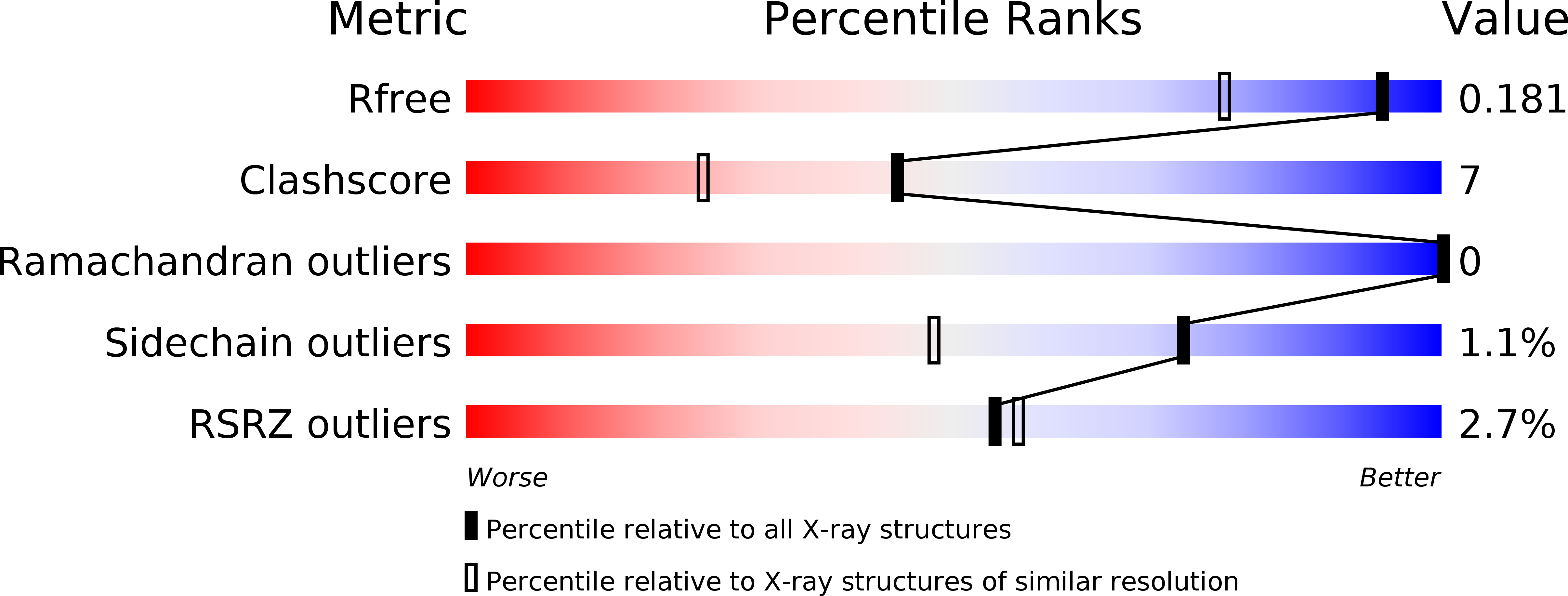
Resolution:
1.45 Å
R-Value Free:
0.18
R-Value Work:
0.16
R-Value Observed:
0.16
Space Group:
P 21 21 21


