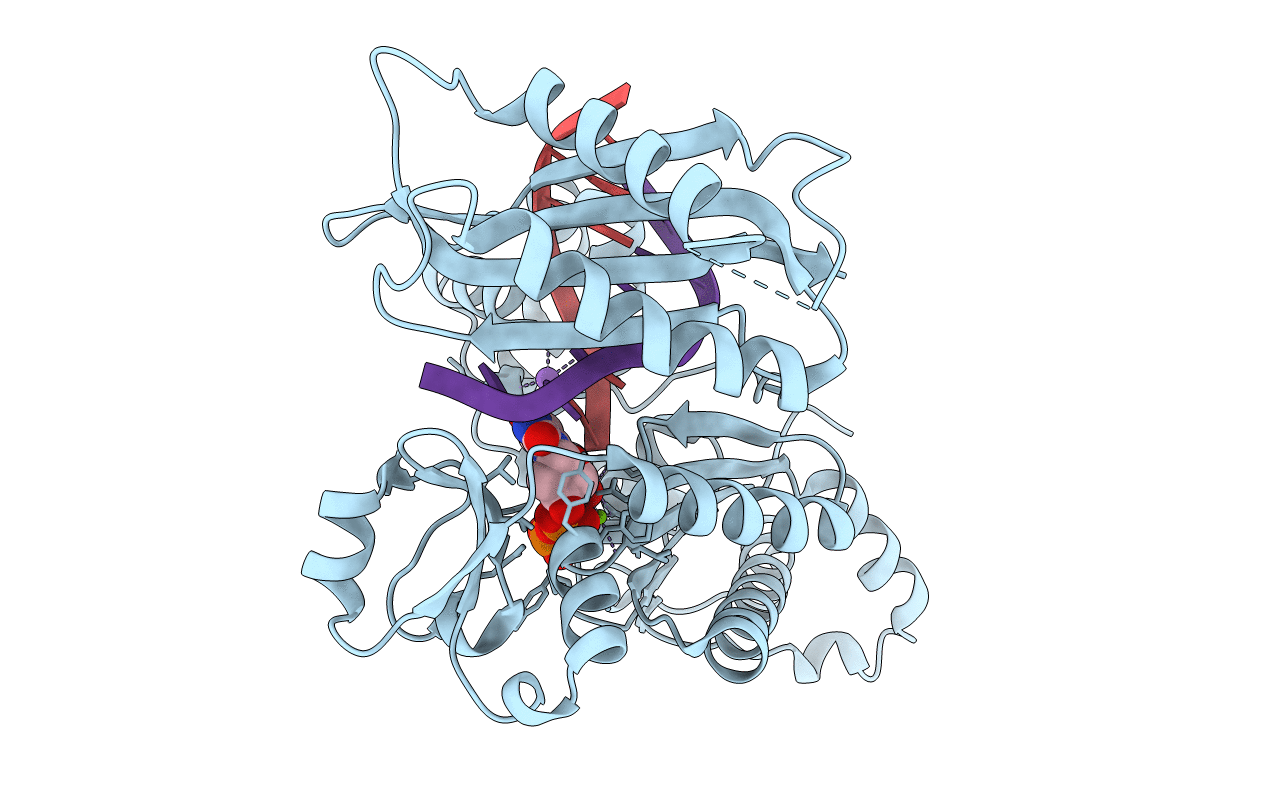
Deposition Date
2010-09-09
Release Date
2010-10-20
Last Version Date
2023-09-06
Entry Detail
PDB ID:
3OSN
Keywords:
Title:
Structural Basis for Proficient Incorporation of dTTP Opposite O6-Methylguanine by Human DNA Polymerase Iota
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.90 Å
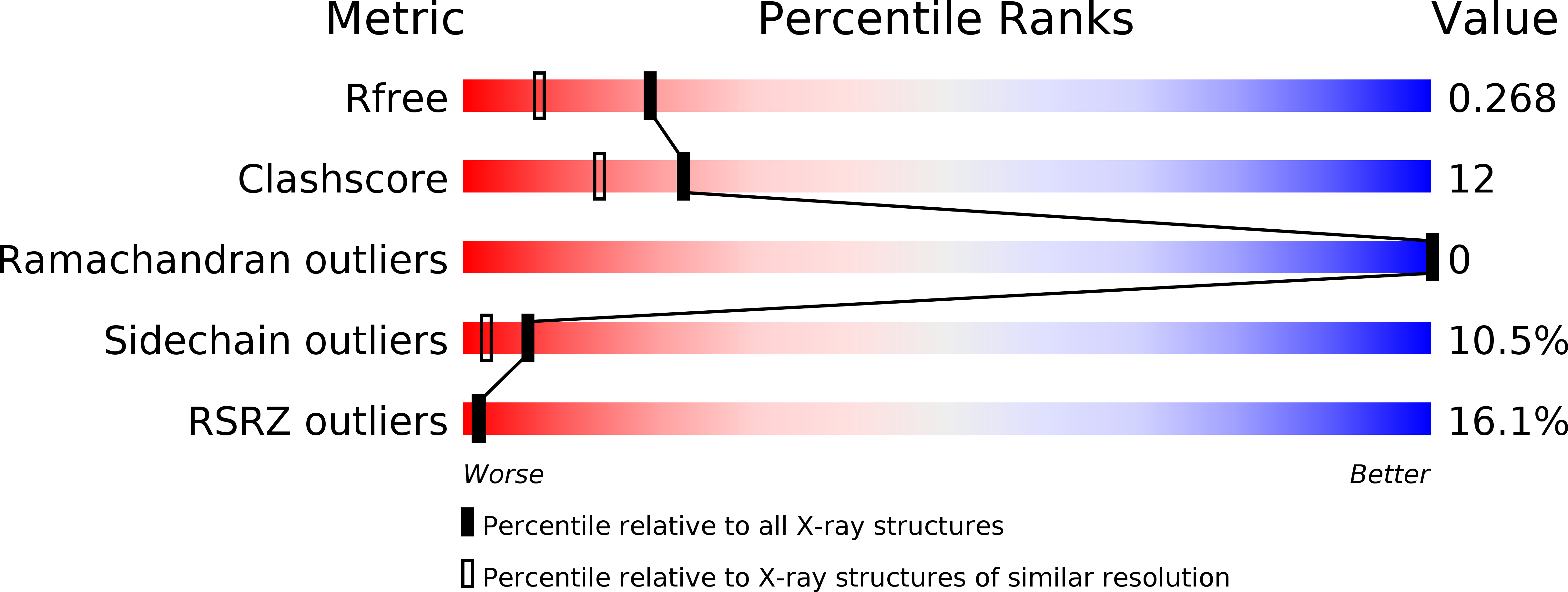
R-Value Free:
0.24
R-Value Work:
0.20
R-Value Observed:
0.20
Space Group:
P 65 2 2