Deposition Date
2010-07-08
Release Date
2011-07-20
Last Version Date
2024-02-21
Entry Detail
PDB ID:
3NVI
Keywords:
Title:
Structure of N-terminal truncated Nop56/58 bound with L7Ae and box C/D RNA
Biological Source:
Source Organism(s):
Pyrococcus furiosus (Taxon ID: 186497)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.71 Å
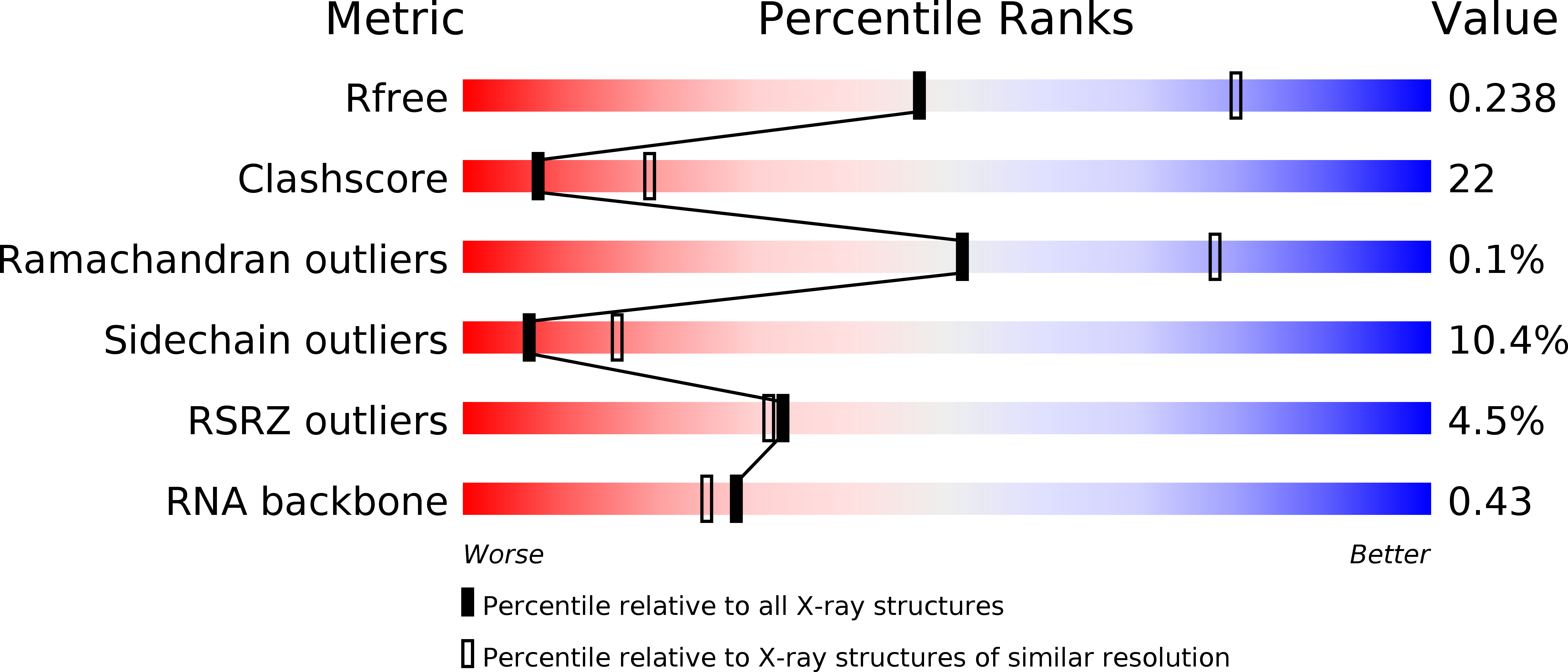
R-Value Free:
0.24
R-Value Work:
0.20
R-Value Observed:
0.21
Space Group:
P 21 21 21