Deposition Date
2009-04-21
Release Date
2009-05-05
Last Version Date
2024-10-30
Entry Detail
PDB ID:
3H50
Keywords:
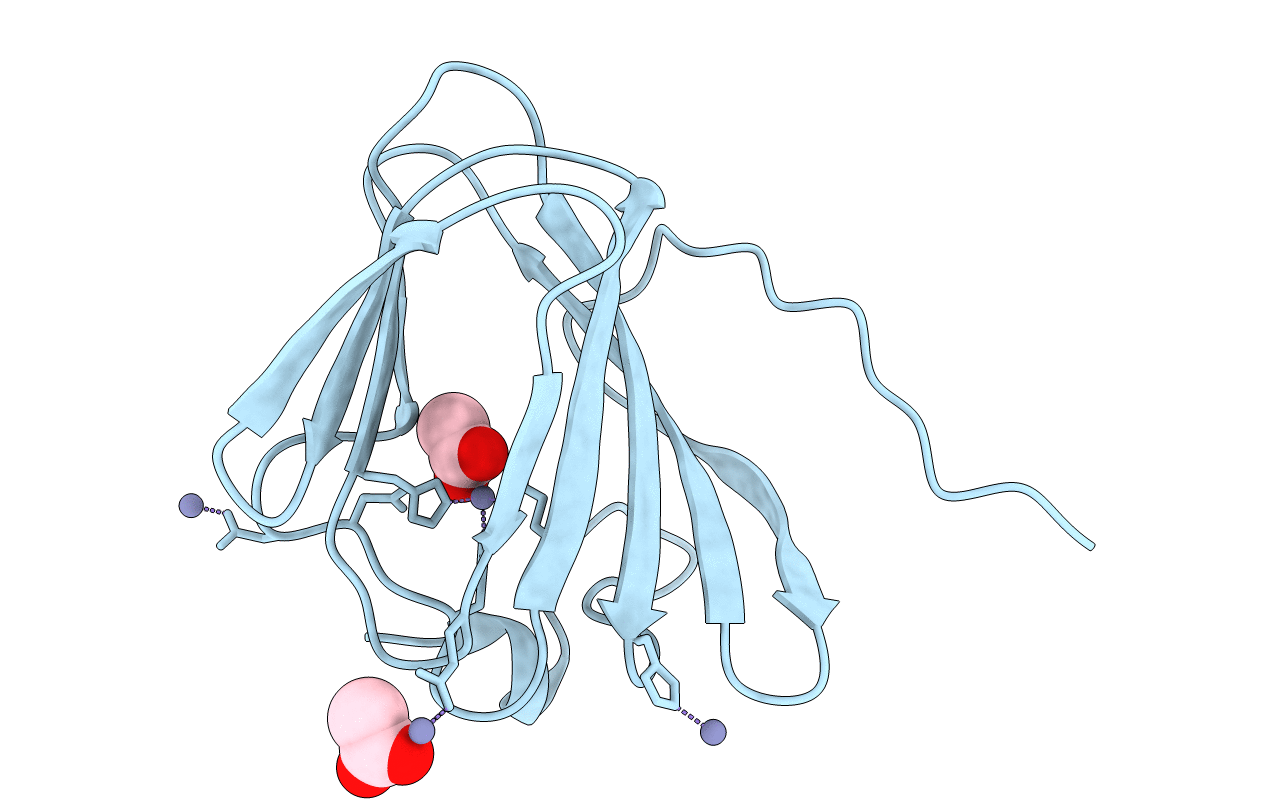
Title:
CRYSTAL STRUCTURE OF A TETRACENOMYCIN POLYKETIDE SYNTHESIS PROTEIN (TCMJ) FROM XANTHOMONAS CAMPESTRIS PV. CAMPESTRIS AT 1.60 A RESOLUTION
Biological Source:
Source Organism(s):
Xanthomonas campestris pv. campestris (Taxon ID: 340)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.60 Å
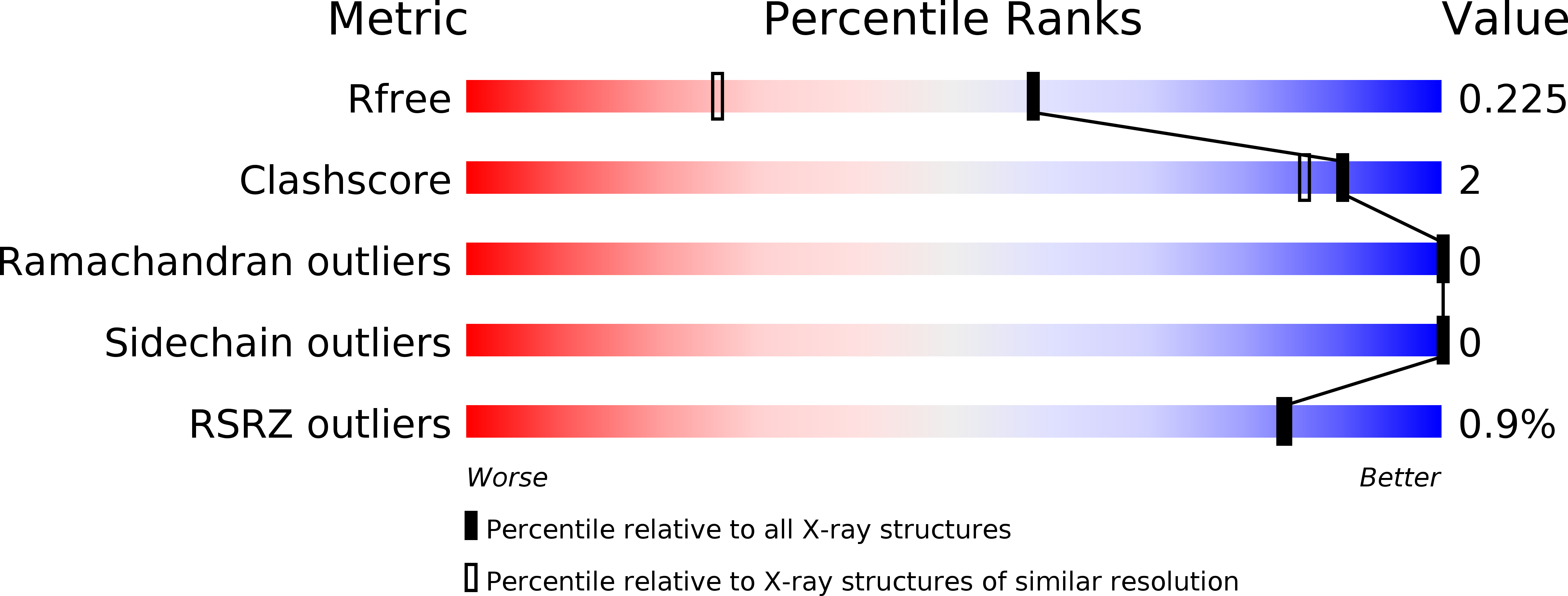
R-Value Free:
0.22
R-Value Work:
0.18
R-Value Observed:
0.18
Space Group:
C 1 2 1