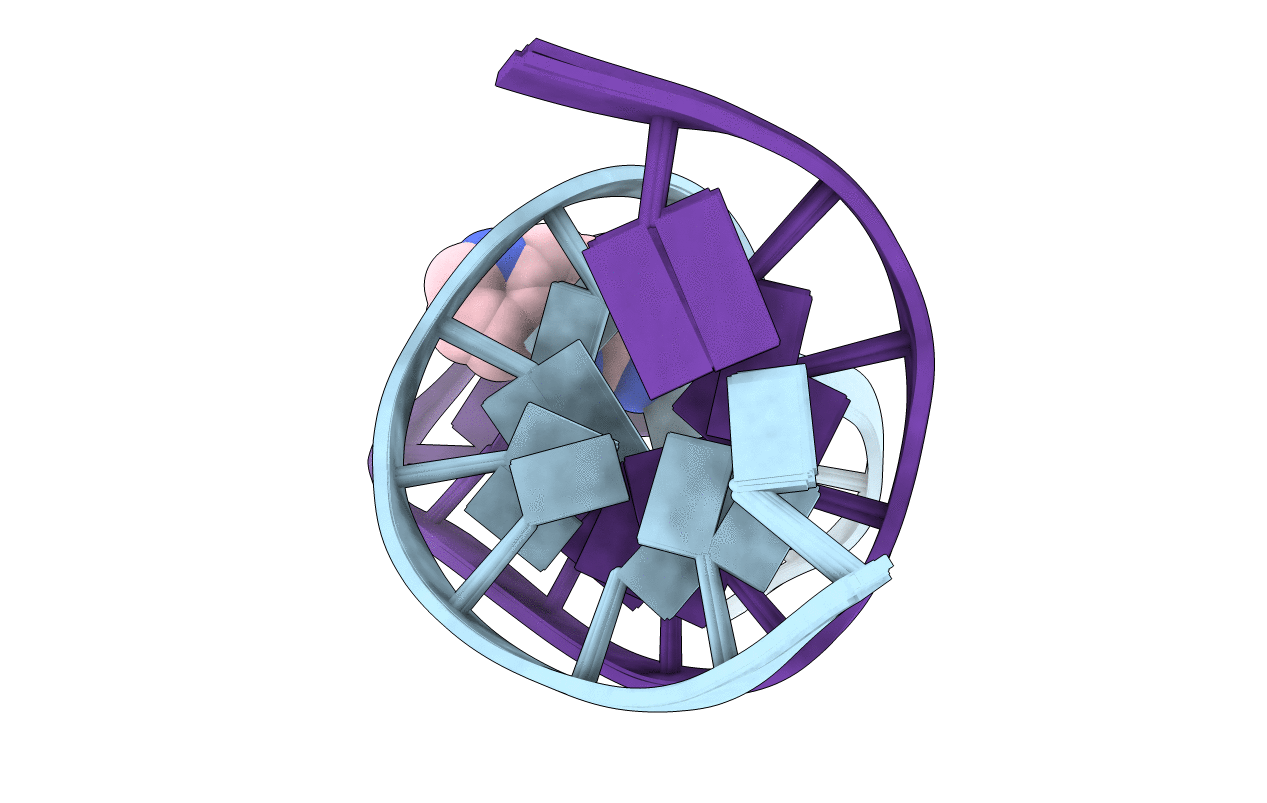Abstact
The conformations of C8-dG adducts of 2-amino-3-methylimidazo[4,5-f]quinoline (IQ) positioned in the C-X1-G, G-X2-C, and C-X3-C contexts in the C-G1-G2-C-G3-C-C recognition sequence of the NarI restriction enzyme were compared, using the oligodeoxynucleotides 5'-d(CTCXGCGCCATC)-3'.5'-d(GATGGCGCCGAG)-3', 5'-d(CTCGXCGCCATC)-3'.5'-d(GATGGCGCCGAG)-3', and 5'-d(CTCGGCXCCATC)-3'.5'-d(GATGGCGCCGAG)-3' (X is the C8-dG adduct of IQ). These were the NarIIQ1, NarIIQ2, and NarIIQ3 duplexes, respectively. In each instance, the glycosyl torsion angle chi for the IQ-modified dG was in the syn conformation. The orientations of the IQ moieties were dependent upon the conformations of torsion angles alpha' [N9-C8-N(IQ)-C2(IQ)] and beta' [C8-N(IQ)-C2(IQ)-N3(IQ)], which were monitored by the patterns of 1H NOEs between the IQ moieties and the DNA in the three sequence contexts. The conformational states of IQ torsion angles alpha' and beta' were predicted from the refined structures of the three adducts obtained from restrained molecular dynamics calculations, utilizing simulated annealing protocols. For the NarIIQ1 and NarIIQ2 duplexes, the alpha' torsion angles were predicted to be -176 +/- 8 degrees and -160 +/- 8 degrees , respectively, whereas for the NarIIQ3 duplex, torsion angle alpha' was predicted to be 159 +/- 7 degrees . Likewise, for the NarIIQ1 and NarIIQ2 duplexes, the beta' torsion angles were predicted to be -152 +/- 8 degrees and -164 +/- 7 degrees , respectively, whereas for the NarIIQ3 duplex, torsion angle beta' was predicted to be -23 +/- 8 degrees . Consequently, the conformations of the IQ adduct in the NarIIQ1 and NarIIQ2 duplexes were similar, with the IQ methyl protons and IQ H4 and H5 protons facing outward in the minor groove, whereas in the NarIIQ3 duplex, the IQ methyl protons and the IQ H4 and H5 protons faced into the DNA duplex, facilitating the base-displaced intercalated orientation of the IQ moiety [Wang, F., Elmquist, C. E., Stover, J. S., Rizzo, C. J., and Stone, M. P. (2006) J. Am. Chem. Soc. 128, 10085-10095]. In contrast, for the NarIIQ1 and NarIIQ2 duplexes, the IQ moiety remained in the minor groove. These sequence-dependent differences suggest that base-displaced intercalation of the IQ adduct is favored when both the 5'- and 3'-flanking nucleotides in the complementary strand are guanines. These conformational differences may correlate with sequence-dependent differences in translesion replication.