Deposition Date
2011-04-05
Release Date
2011-07-06
Last Version Date
2024-10-16
Entry Detail
PDB ID:
2YFD
Keywords:
Title:
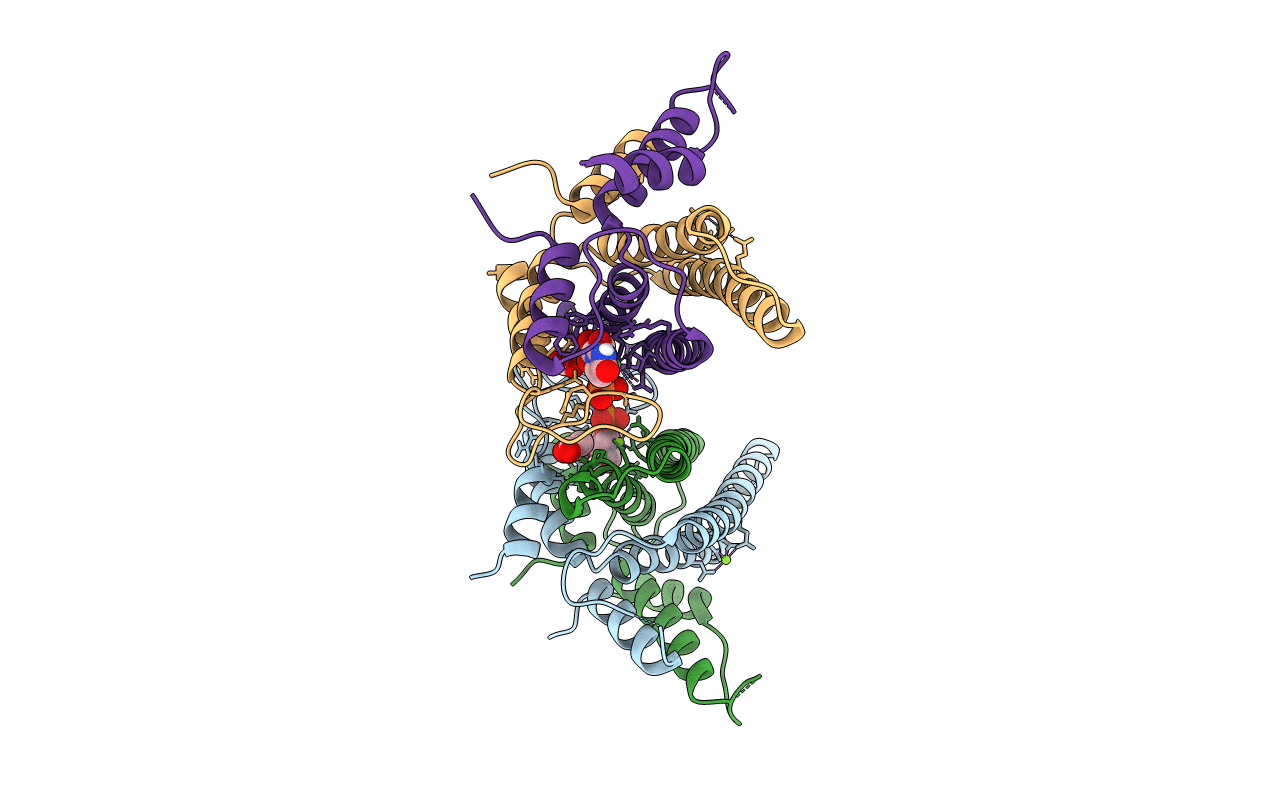
STRUCTURAL AND FUNCTIONAL INSIGHTS OF DR2231 PROTEIN, THE MAZG-LIKE NUCLEOSIDE TRIPHOSPHATE PYROPHOSPHOHYDROLASE FROM DEINOCOCCUS RADIODURANS, COMPLEXED WITH Mg and dUMP
Biological Source:
Source Organism(s):
DEINOCOCCUS RADIODURANS (Taxon ID: 243230)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.77 Å
R-Value Free:
0.21
R-Value Work:
0.18
R-Value Observed:
0.18
Space Group:
P 21 21 2


