Deposition Date
2011-01-20
Release Date
2011-06-08
Last Version Date
2023-12-20
Entry Detail
PDB ID:
2Y68
Keywords:
Title:
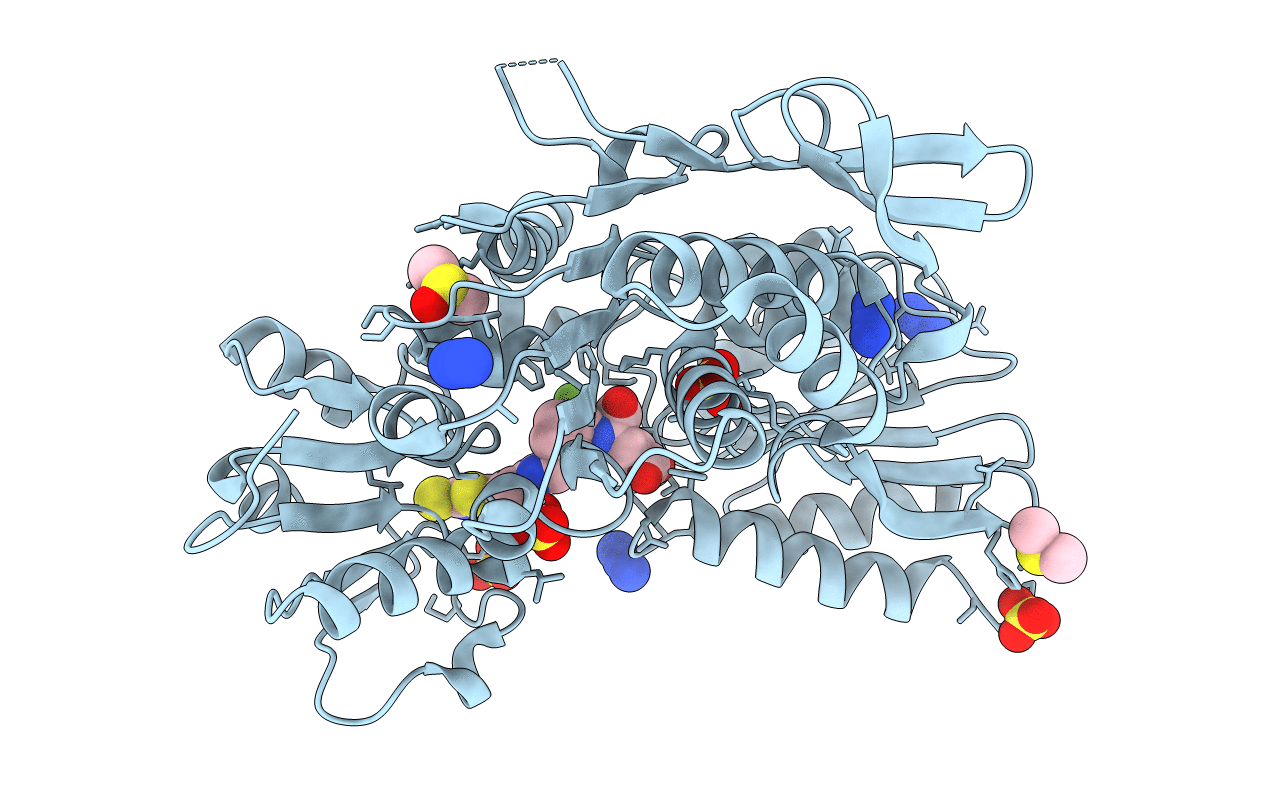
Structure-based design of a new series of D-glutamic acid-based inhibitors of bacterial MurD ligase
Biological Source:
Source Organism(s):
Escherichia coli K-12 (Taxon ID: 83333)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.49 Å
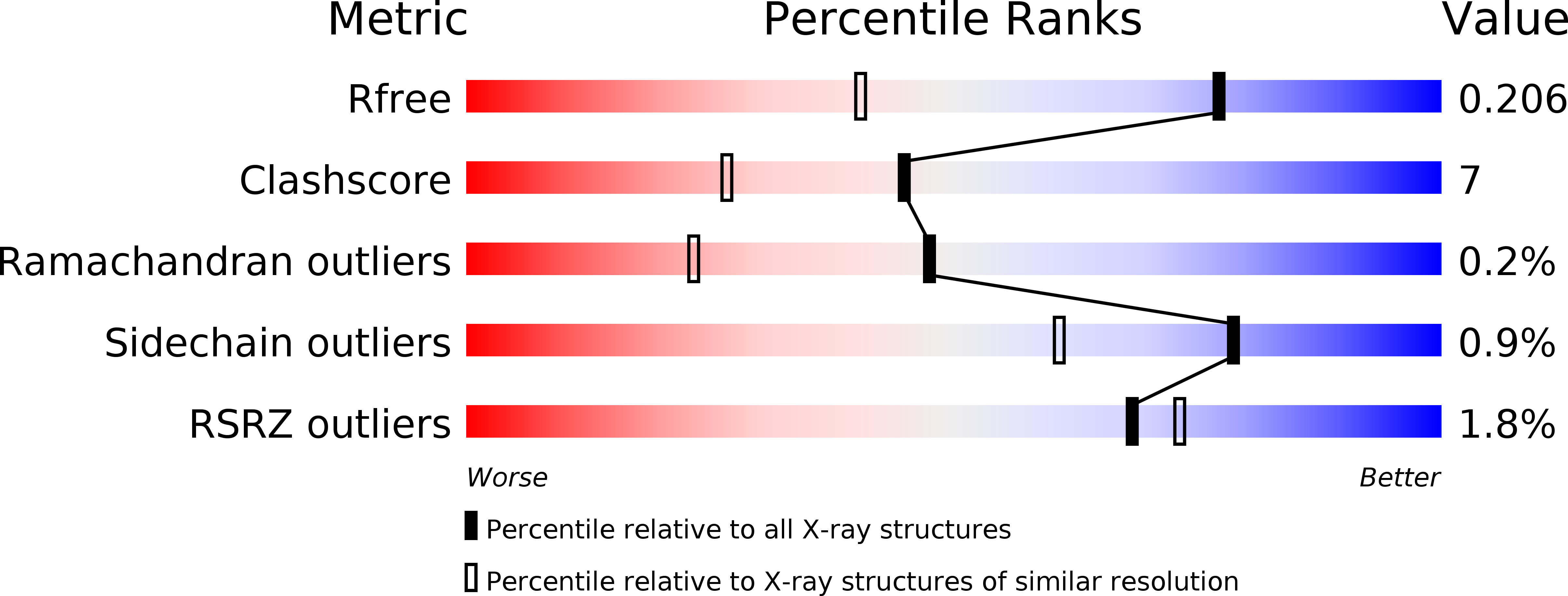
R-Value Free:
0.20
R-Value Work:
0.17
R-Value Observed:
0.18
Space Group:
P 41