Deposition Date
2009-11-20
Release Date
2010-08-25
Last Version Date
2024-11-13
Entry Detail
PDB ID:
2WYS
Keywords:
Title:
High resolution crystallographic structure of the Clostridium thermocellum N-terminal endo-1,4-beta-D-xylanase 10B (Xyn10B) CBM22-1- GH10 modules complexed with xylohexaose
Biological Source:
Source Organism(s):
CLOSTRIDIUM THERMOCELLUM (Taxon ID: 1515)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.75 Å
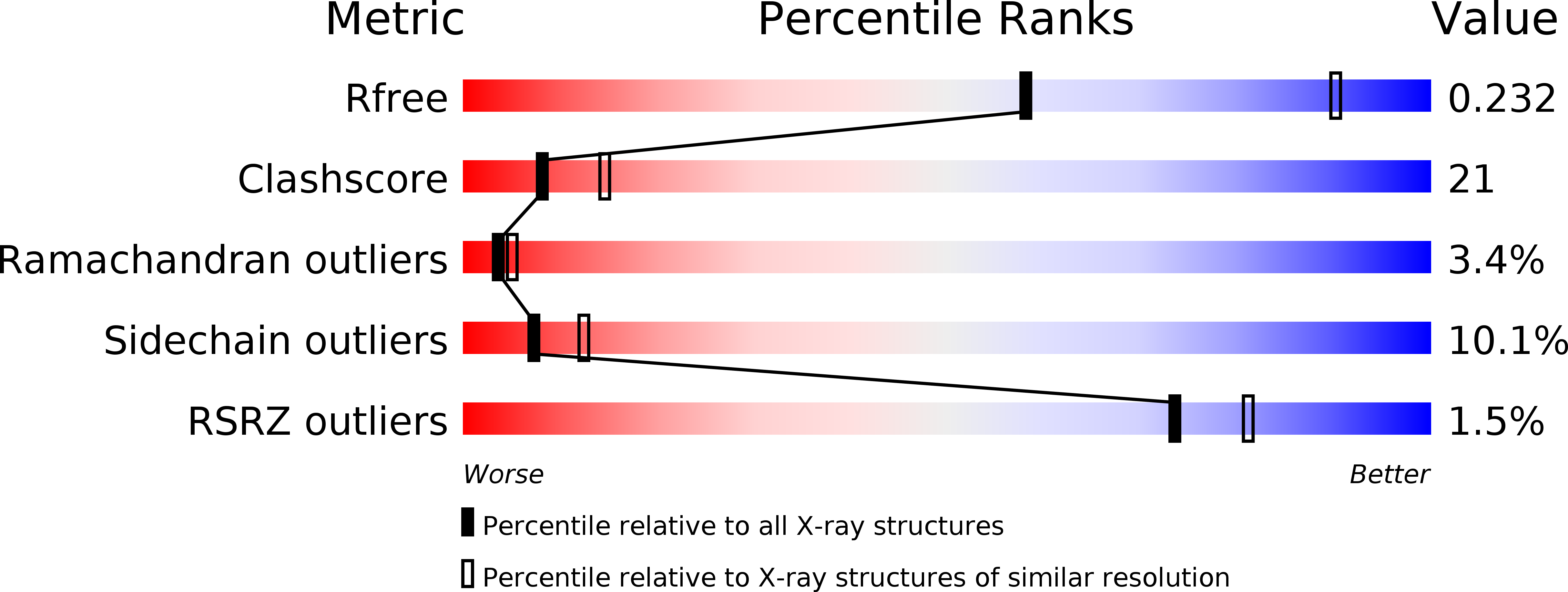
R-Value Free:
0.23
R-Value Work:
0.18
R-Value Observed:
0.18
Space Group:
P 32 2 1