Deposition Date
2007-06-15
Release Date
2007-12-04
Last Version Date
2023-08-30
Entry Detail
PDB ID:
2QB2
Keywords:
Title:
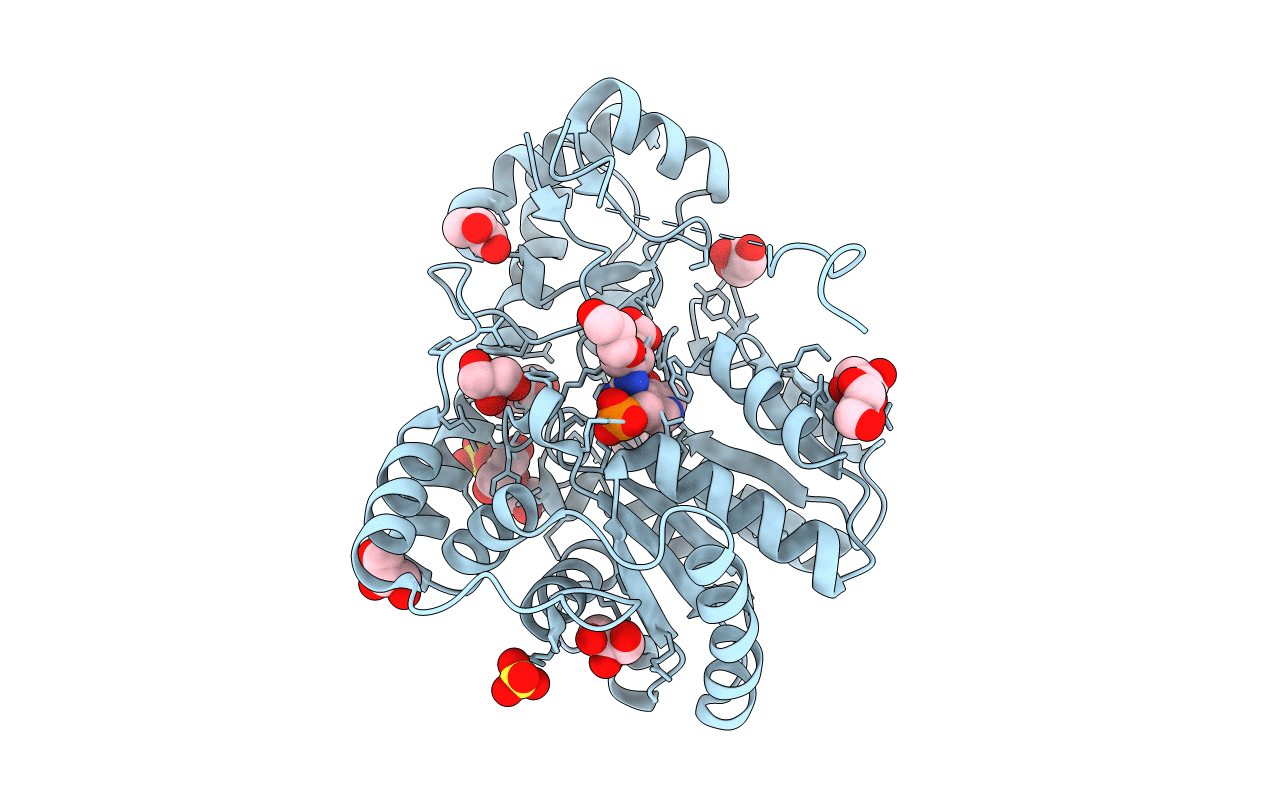
Structural Studies Reveal the Inactivation of E. coli L-aspartate aminotransferase by (s)-4,5-dihydro-2thiophenecarboylic acid (SADTA) via two mechanisms (at pH 7.0).
Biological Source:
Source Organism(s):
Escherichia coli (Taxon ID: 562)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.70 Å
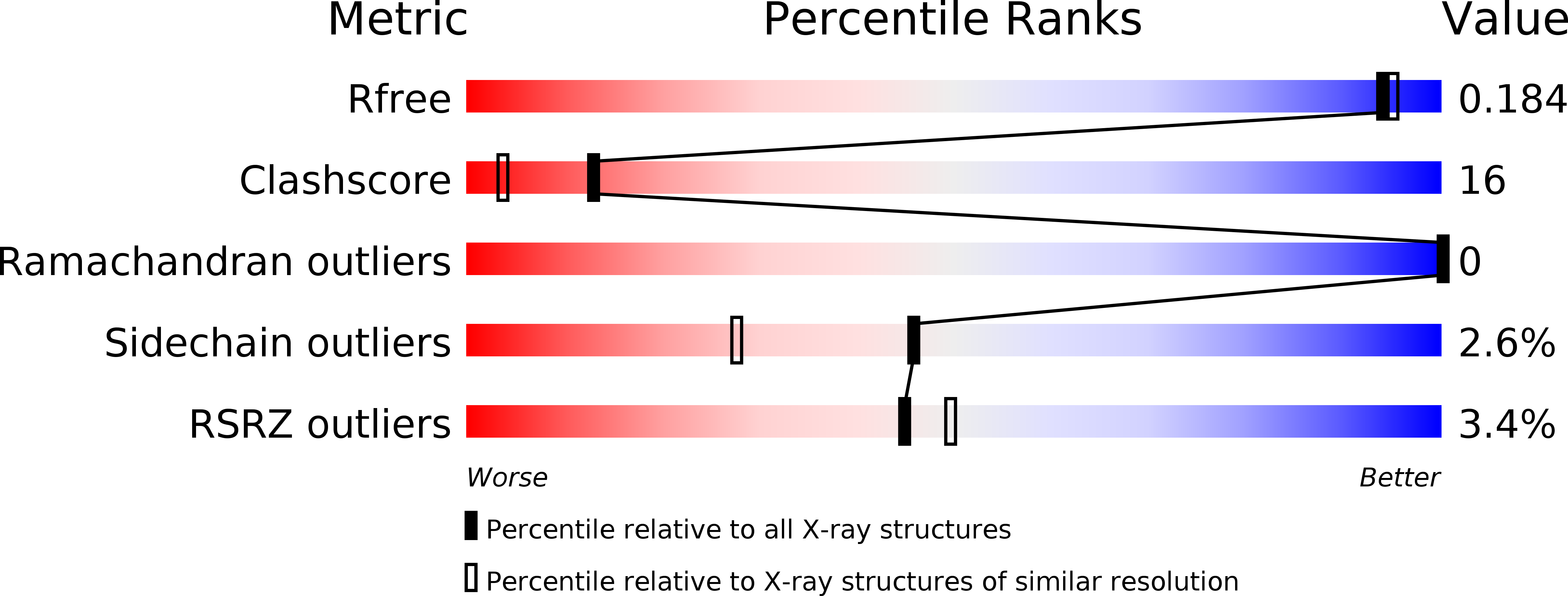
R-Value Free:
0.18
R-Value Work:
0.14
R-Value Observed:
0.14
Space Group:
C 2 2 21