Deposition Date
2012-06-19
Release Date
2012-07-04
Last Version Date
2024-05-15
Entry Detail
PDB ID:
2LUP
Keywords:
Title:
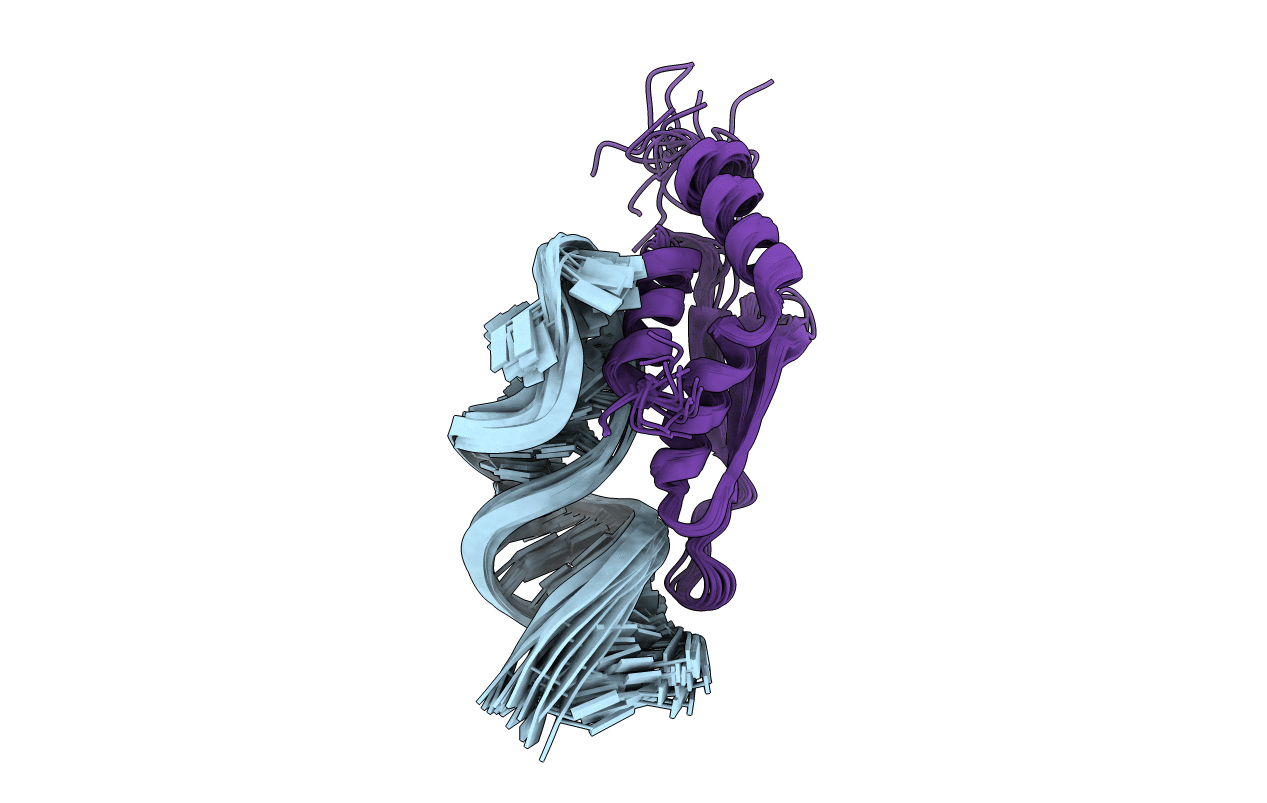
RDC refined solution structure of double-stranded RNA binding domain of S. cerevisiae RNase III (rnt1p) in complex with the terminal RNA hairpin of snr47 precursor
Biological Source:
Source Organism(s):
Saccharomyces cerevisiae (Taxon ID: 559292)
Expression System(s):
Method Details:
Experimental Method:
Conformers Calculated:
100
Conformers Submitted:
16
Selection Criteria:
structures with the lowest energy