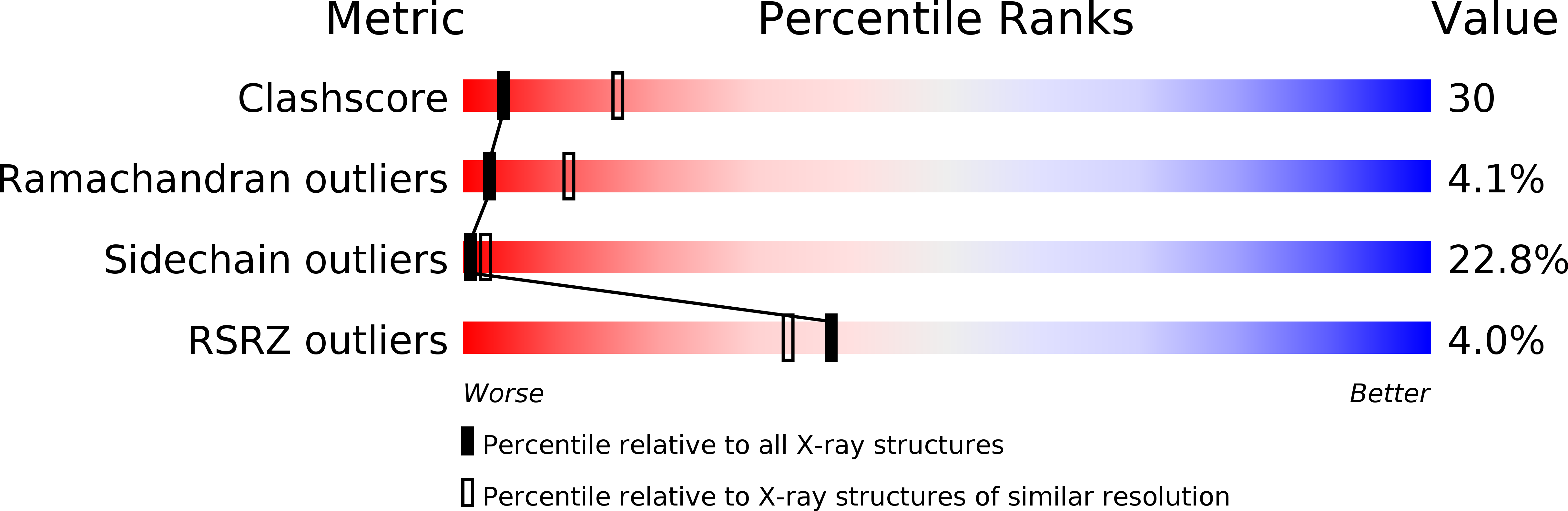
Deposition Date
2006-09-18
Release Date
2007-01-30
Last Version Date
2024-10-16
Entry Detail
PDB ID:
2J5L
Keywords:
Title:
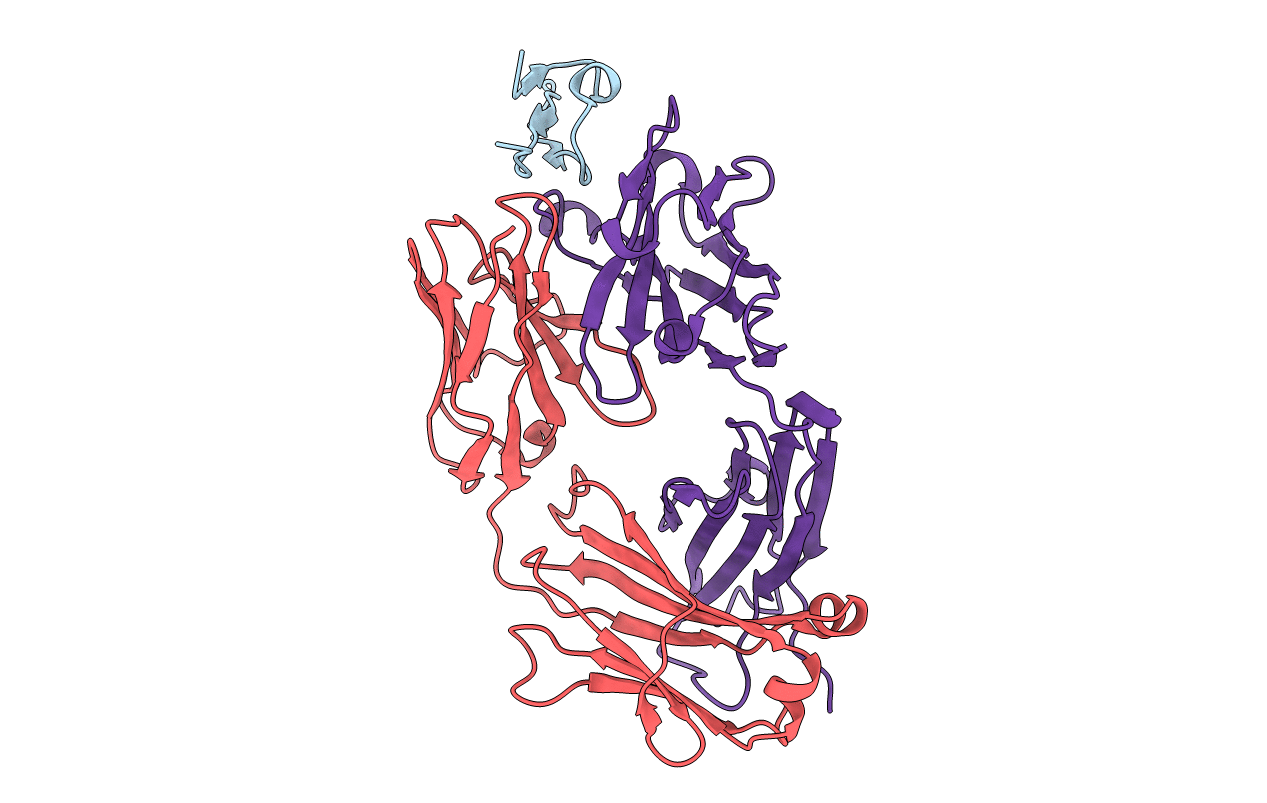
Structure of a Plasmodium falciparum apical membrane antigen 1-Fab F8. 12.19 complex
Biological Source:
Source Organism(s):
PLASMODIUM FALCIPARUM (Taxon ID: 5833)
MUS MUSCULUS (Taxon ID: 10090)
MUS MUSCULUS (Taxon ID: 10090)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.90 Å
R-Value Free:
0.27
R-Value Work:
0.21
R-Value Observed:
0.21
Space Group:
P 63