Deposition Date
1995-11-13
Release Date
1996-06-10
Last Version Date
2024-10-09
Entry Detail
PDB ID:
2BTB
Keywords:
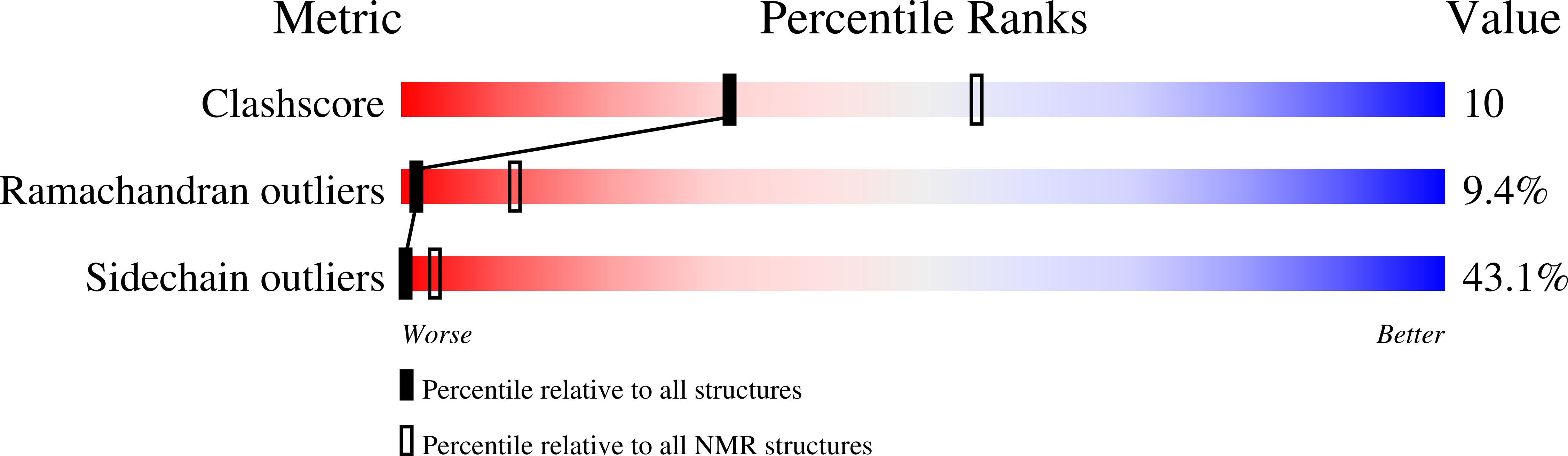
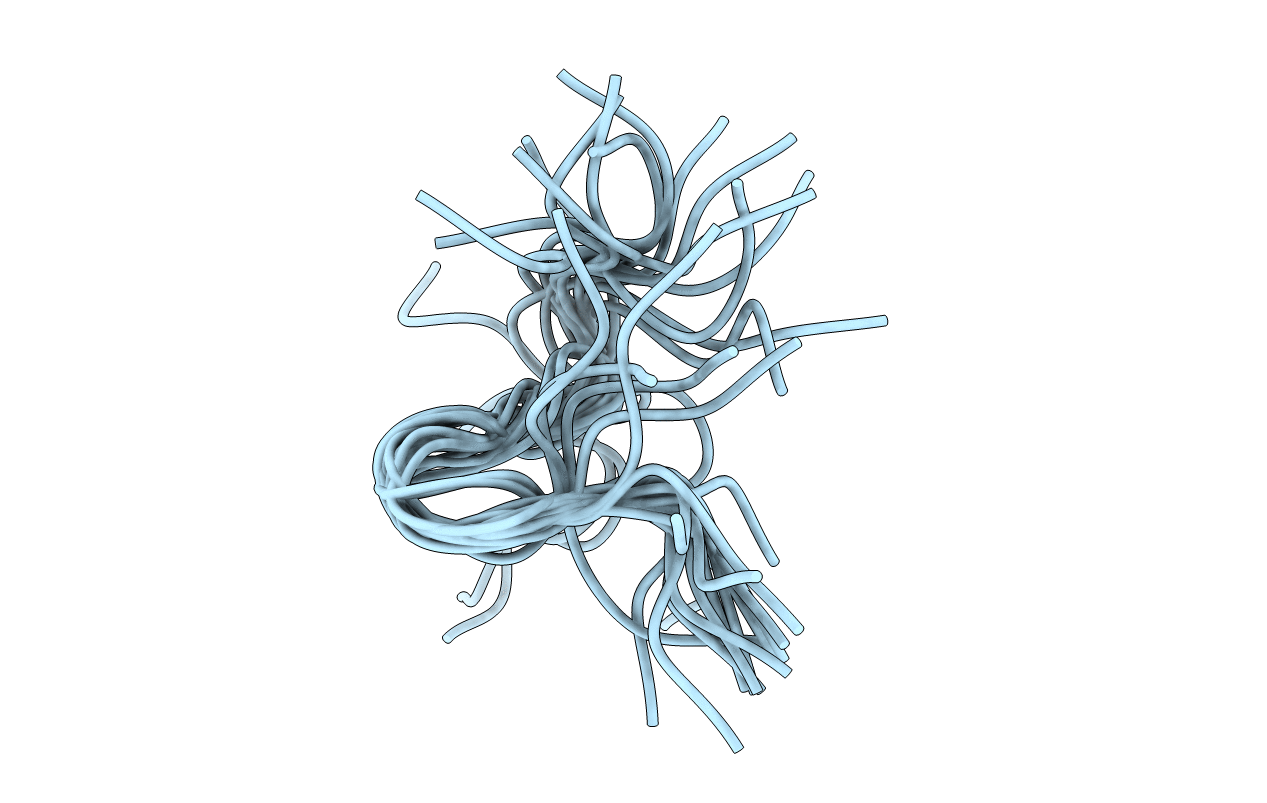
Title:
NMR STUDY OF N-TERMINAL HUMAN BAND 3 PEPTIDE, RESIDUES 1-15
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Method Details:
Experimental Method:
Conformers Submitted:
20