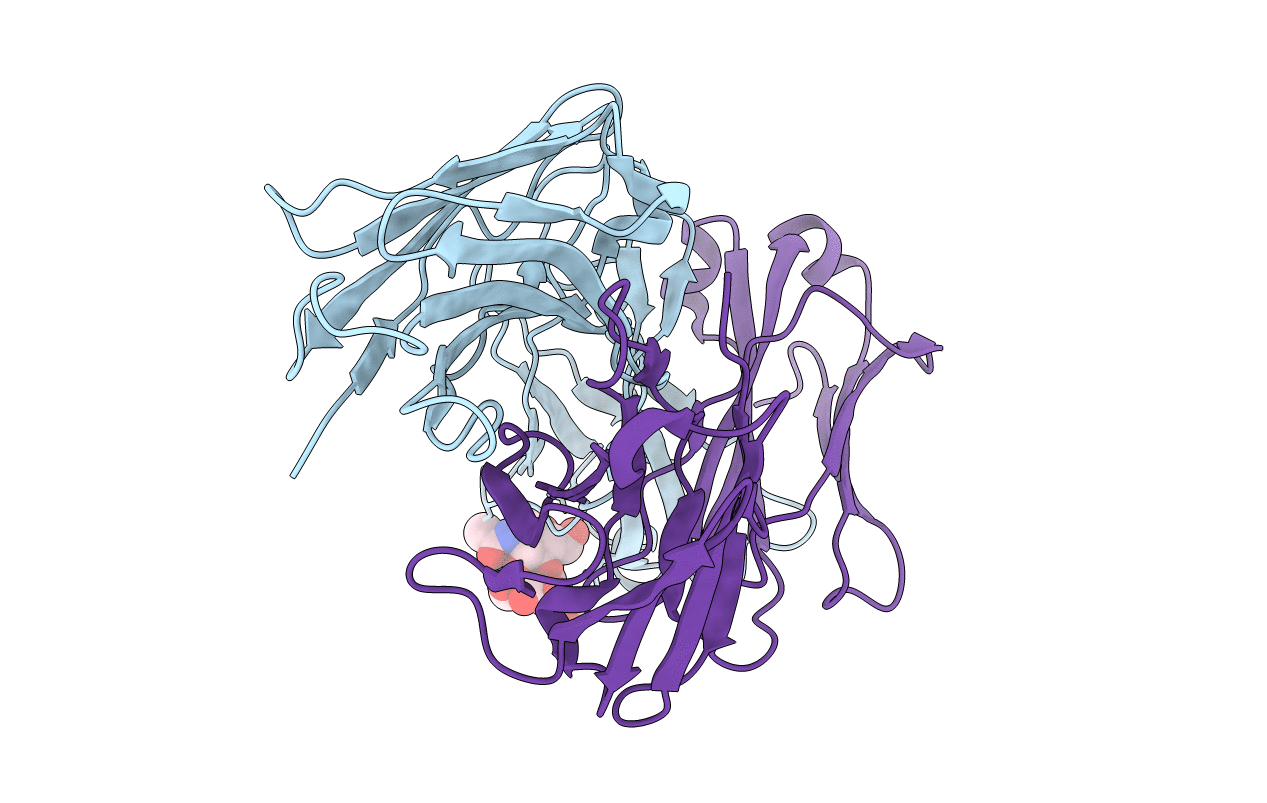Abstact
The 2.6 A (1 A=0.1 nm) resolution structure has been determined for the glycosylated Fab (fragment antigen binding) of an IgM (Yvo) obtained from a subject with Waldenström's macroglobulinaemia. Dynamic light scattering was used to estimate the gel point and monitor the formation of an ordered hydroscopic gel of Yvo IgM upon cooling. If a cryoglobulin forms gels in peripheral tissues and organs, the associated swelling and damage to microvasculature can result in considerable morbidity and mortality. The three-dimensional structure of the branched N-linked oligosaccharide associated with the CH1 domain (first constant domain of heavy chain) is reported. The carbohydrate may act to shield part of the lateral surface of the CH1 domain and crowd the junction between the CH1 and CH2 domains, thereby limiting the segmental flexibility of the Fab arms in intact Yvo IgM, especially at low temperatures. Recently, Yvo IgM was shown to have the properties of a naturally occurring proteolytic antibody [Paul, Karle, Planque, Taguchi, Salas, Nishiyama, Handy, Hunter, Edmundson and Hanson (2004) J. Biol. Chem. 279, 39611-39619; Planque, Bangale, Song, Karle, Taguchi, Poindexter, Bick, Edmundson, Nishiyama and Paul (2004) J. Biol Chem. 279, 14024-14032]. The Yvo protein displayed the ability to cleave, by a nucleophilic mechanism, the amide bonds of a variety of serine protease substrates and the gp120 coat protein of HIV. An atypical serine, arginine and glutamate motif is located in the middle of the Yvo antigen-binding site and displays an overall geometry that mimics the classical serine, histidine and aspartate catalytic triad of serine proteases. Our present findings indicate that pre-existing or natural antibodies can utilize at least one novel strategy for the cleavage of peptide bonds.