Deposition Date
2005-06-30
Release Date
2005-10-04
Last Version Date
2025-03-26
Entry Detail
PDB ID:
2A5H
Keywords:
Title:
2.1 Angstrom X-ray crystal structure of lysine-2,3-aminomutase from Clostridium subterminale SB4, with Michaelis analog (L-alpha-lysine external aldimine form of pyridoxal-5'-phosphate).
Biological Source:
Source Organism(s):
Clostridium subterminale (Taxon ID: 1550)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.10 Å
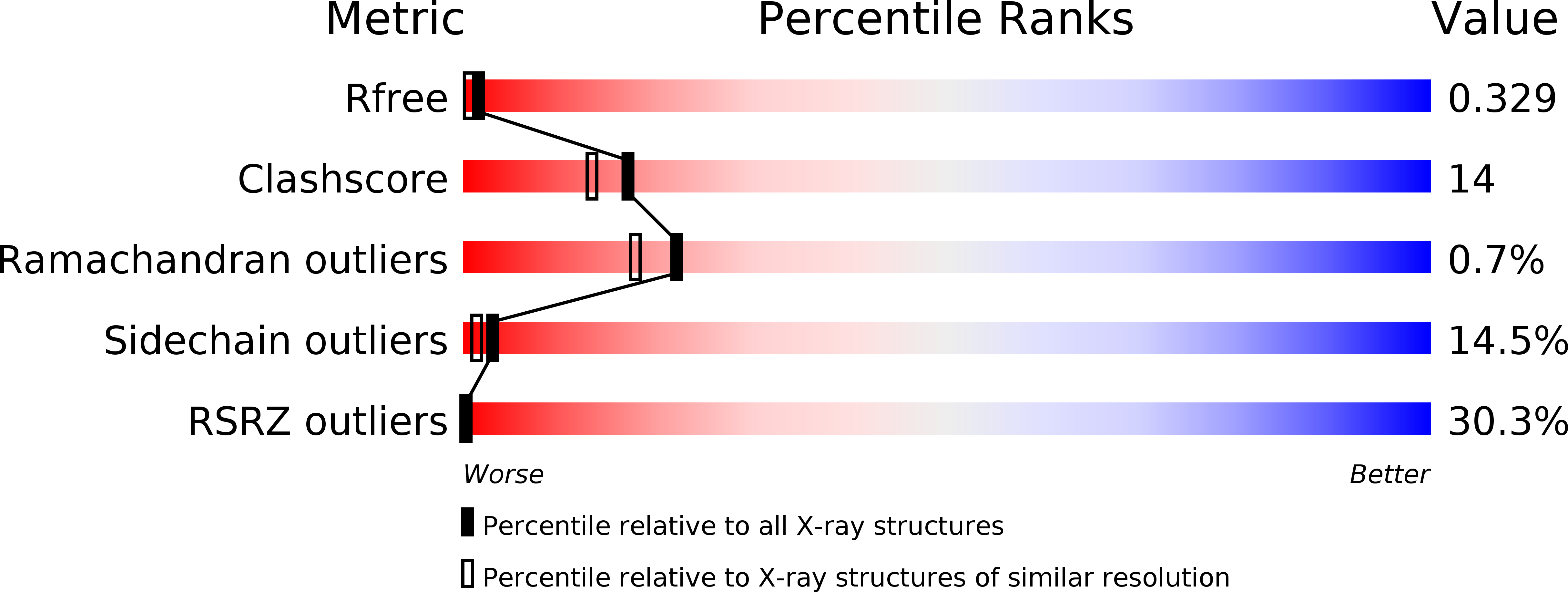
R-Value Free:
0.22
R-Value Work:
0.18
R-Value Observed:
0.18
Space Group:
C 1 2 1


