Deposition Date
2004-11-19
Release Date
2005-01-19
Last Version Date
2024-05-15
Entry Detail
PDB ID:
1WCR
Keywords:
Title:
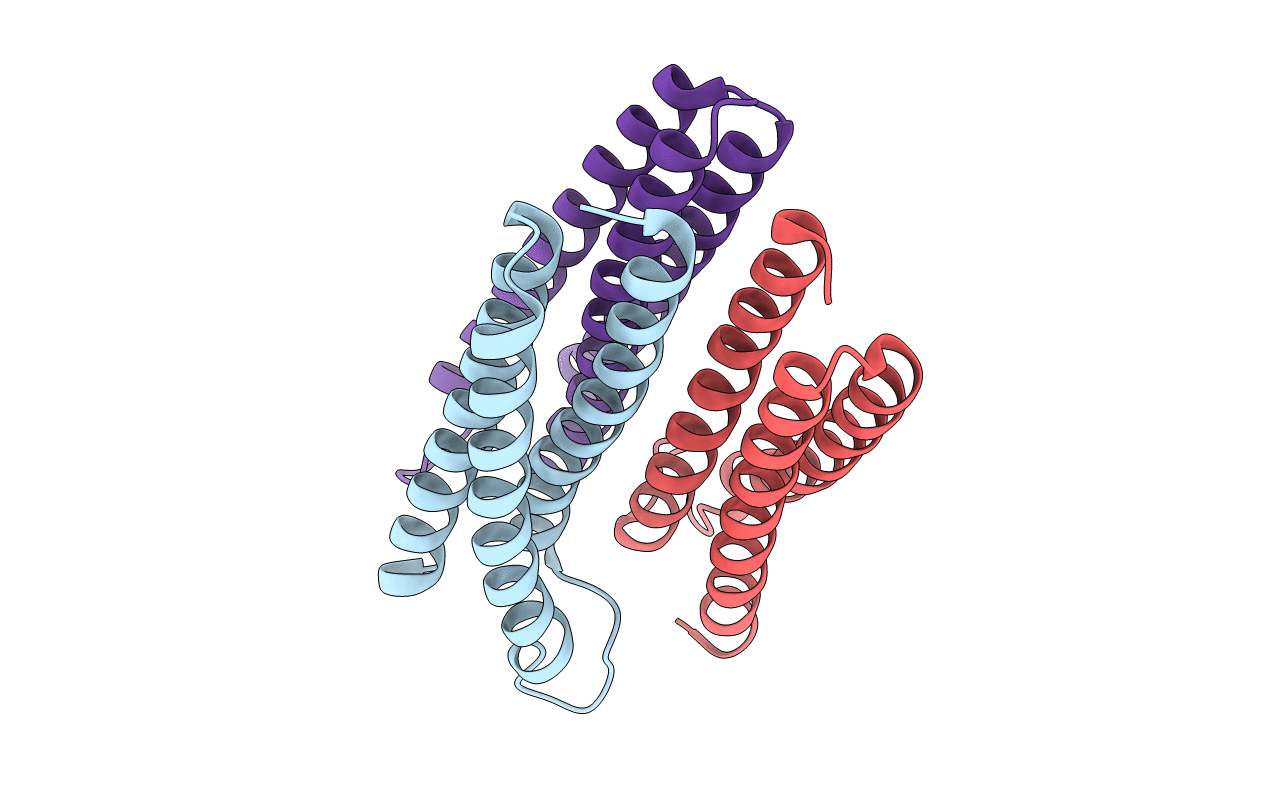
Trimeric Structure of the Enzyme IIA from Escherichia coli Phosphotransferase System Specific for N,N'-Diacetylchitobiose
Biological Source:
Source Organism(s):
ESCHERICHIA COLI (Taxon ID: 562)
Expression System(s):
Method Details:
Experimental Method:
Conformers Calculated:
100
Conformers Submitted:
1
Selection Criteria:
RESTRAINED REGULARIZED MEAN STRUCTURE